Increase of enzyme activity through specific covalent modification with fragments
- PMID: 29163914
- PMCID: PMC5674511
- DOI: 10.1039/c7sc01966a
Increase of enzyme activity through specific covalent modification with fragments
Abstract
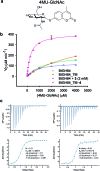
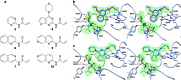
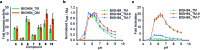
Modulation of enzyme activity is a powerful means of probing cellular function and can be exploited for diverse applications. Here, we explore a method of enzyme activation where covalent tethering of a small molecule to an enzyme can increase catalytic activity (kcat/KM) up to 35-fold. Using a bacterial glycoside hydrolase, BtGH84, we demonstrate how small molecule "fragments", identified as activators in free solution, can be covalently tethered to the protein using Michael-addition chemistry. We show how tethering generates a constitutively-activated enzyme-fragment conjugate, which displays both improved catalytic efficiency and increased susceptibility to certain inhibitor classes. Structure guided modifications of the tethered fragment demonstrate how specific interactions between the fragment and the enzyme influence the extent of activation. This work suggests that a similar approach may be used to modulate the activity of enzymes such as to improve catalytic efficiency or increase inhibitor susceptibility.
Figures
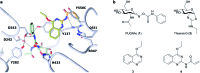
Similar articles
-
Fragment-derived modulators of an industrial β-glucosidase.Biochem J. 2020 Nov 27;477(22):4383-4395. doi: 10.1042/BCJ20200507. Biochem J. 2020. PMID: 33111951 Free PMC article.
-
Discovery of selective small-molecule activators of a bacterial glycoside hydrolase.Angew Chem Int Ed Engl. 2014 Dec 1;53(49):13419-23. doi: 10.1002/anie.201407081. Epub 2014 Oct 7. Angew Chem Int Ed Engl. 2014. PMID: 25291993 Free PMC article.
-
Covalent Tethering of Fragments For Covalent Probe Discovery.Medchemcomm. 2016 Apr 1;7(4):576-585. doi: 10.1039/c5md00518c. Epub 2016 Jan 28. Medchemcomm. 2016. PMID: 27398190 Free PMC article.
-
Probing structural adaptivity at PPI interfaces with small molecules.Drug Discov Today Technol. 2013 Dec;10(4):e501-8. doi: 10.1016/j.ddtec.2012.10.009. Drug Discov Today Technol. 2013. PMID: 24451641 Review.
-
Biophysical and computational fragment-based approaches to targeting protein-protein interactions: applications in structure-guided drug discovery.Q Rev Biophys. 2012 Nov;45(4):383-426. doi: 10.1017/S0033583512000108. Epub 2012 Sep 13. Q Rev Biophys. 2012. PMID: 22971516 Review.
Cited by
-
Atrazine Degradation Using Immobilized Triazine Hydrolase from Arthrobacter aurescens TC1 in Mesoporous Silica Nanomaterials.ACS Environ Au. 2023 Nov 1;3(6):361-369. doi: 10.1021/acsenvironau.3c00036. eCollection 2023 Nov 15. ACS Environ Au. 2023. PMID: 38028742 Free PMC article.
-
Fragment-derived modulators of an industrial β-glucosidase.Biochem J. 2020 Nov 27;477(22):4383-4395. doi: 10.1042/BCJ20200507. Biochem J. 2020. PMID: 33111951 Free PMC article.
-
Role of rare-earth elements in enhancing bioelectrocatalysis for biosensing with NAD+-dependent glutamate dehydrogenase.Chem Sci. 2021 Sep 9;12(40):13434-13441. doi: 10.1039/d1sc00193k. eCollection 2021 Oct 20. Chem Sci. 2021. PMID: 34777762 Free PMC article.
-
Identification of Cysteine 270 as a Novel Site for Allosteric Modulators of SARS-CoV-2 Papain-Like Protease.Angew Chem Int Ed Engl. 2022 Dec 23;61(52):e202212378. doi: 10.1002/anie.202212378. Epub 2022 Nov 28. Angew Chem Int Ed Engl. 2022. PMID: 36308706 Free PMC article.
-
Modulated Protein Binding Ability of Anti-Diabetic Drugs in Presence of Monodispersed Gold Nanoparticles and its Inhibitory Potential towards Advanced Glycated End (AGE) Product Formation.J Fluoresc. 2020 Jan;30(1):193-204. doi: 10.1007/s10895-019-02485-y. Epub 2020 Jan 10. J Fluoresc. 2020. PMID: 31925653
References
-
- Nurse P., Thuriaux P., Nasmyth K. Mol. Gen. Genet. 1976;146:167–178. - PubMed
-
- Panaretou B., Siligardi G., Meyer P., Maloney A., Sullivan J. K., Singh S., Millson S. H., Clarke P. A., Naaby-Hansen S., Stein R., Cramer R., Mollapour M., Workman P., Piper P. W., Pearl L. H., Prodromou C. Mol. Cell. 2002;10:1307–1318. - PubMed
-
- Scheffzek K., Ahmadian M. R., Kabsch W., Wiesmuller L., Lautwein A., Schmitz F., Wittinghofer A. Science. 1997;277:333–338. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

