Protein complex scaffolding predicted as a prevalent function of long non-coding RNAs
- PMID: 29165713
- PMCID: PMC5778612
- DOI: 10.1093/nar/gkx1169
Protein complex scaffolding predicted as a prevalent function of long non-coding RNAs
Abstract
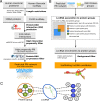
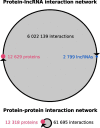
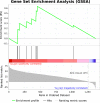
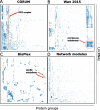
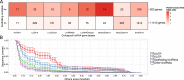
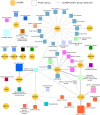
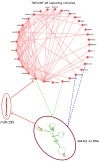
The human transcriptome contains thousands of long non-coding RNAs (lncRNAs). Characterizing their function is a current challenge. An emerging concept is that lncRNAs serve as protein scaffolds, forming ribonucleoproteins and bringing proteins in proximity. However, only few scaffolding lncRNAs have been characterized and the prevalence of this function is unknown. Here, we propose the first computational approach aimed at predicting scaffolding lncRNAs at large scale. We predicted the largest human lncRNA-protein interaction network to date using the catRAPID omics algorithm. In combination with tissue expression and statistical approaches, we identified 847 lncRNAs (∼5% of the long non-coding transcriptome) predicted to scaffold half of the known protein complexes and network modules. Lastly, we show that the association of certain lncRNAs to disease may involve their scaffolding ability. Overall, our results suggest for the first time that RNA-mediated scaffolding of protein complexes and modules may be a common mechanism in human cells.
© The Author(s) 2017. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

