Regulation of G2/M Transition by Inhibition of WEE1 and PKMYT1 Kinases
- PMID: 29168755
- PMCID: PMC6149964
- DOI: 10.3390/molecules22122045
Regulation of G2/M Transition by Inhibition of WEE1 and PKMYT1 Kinases
Abstract
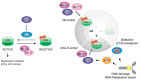
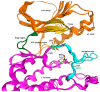
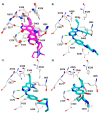
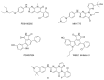
In the cell cycle, there are two checkpoint arrests that allow cells to repair damaged DNA in order to maintain genomic integrity. Many cancer cells have defective G1 checkpoint mechanisms, thus depending on the G2 checkpoint far more than normal cells. G2 checkpoint abrogation is therefore a promising concept to preferably damage cancerous cells over normal cells. The main factor influencing the decision to enter mitosis is a complex composed of Cdk1 and cyclin B. Cdk1/CycB is regulated by various feedback mechanisms, in particular inhibitory phosphorylations at Thr14 and Tyr15 of Cdk1. In fact, Cdk1/CycB activity is restricted by the balance between WEE family kinases and Cdc25 phosphatases. The WEE kinase family consists of three proteins: WEE1, PKMYT1, and the less important WEE1B. WEE1 exclusively mediates phosphorylation at Tyr15, whereas PKMYT1 is dual-specific for Tyr15 as well as Thr14. Inhibition by a small molecule inhibitor is therefore proposed to be a promising option since WEE kinases bind Cdk1, altering equilibria and thus affecting G2/M transition.
Keywords: G2/M transition; PKMYT1; WEE1.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


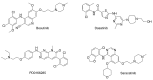
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

