Complexity and conservation of regulatory landscapes underlie evolutionary resilience of mammalian gene expression
- PMID: 29180706
- PMCID: PMC5733139
- DOI: 10.1038/s41559-017-0377-2
Complexity and conservation of regulatory landscapes underlie evolutionary resilience of mammalian gene expression
Abstract
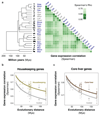
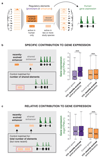
To gain insight into how mammalian gene expression is controlled by rapidly evolving regulatory elements, we jointly analysed promoter and enhancer activity with downstream transcription levels in liver samples from 15 species. Genes associated with complex regulatory landscapes generally exhibit high expression levels that remain evolutionarily stable. While the number of regulatory elements is the key driver of transcriptional output and resilience, regulatory conservation matters: elements active across mammals most effectively stabilize gene expression. In contrast, recently evolved enhancers typically contribute weakly, consistent with their high evolutionary plasticity. These effects are observed across the entire mammalian clade and are robust to potential confounders, such as the gene expression level. Using liver as a representative somatic tissue, our results illuminate how the evolutionary stability of gene expression is profoundly entwined with both the number and conservation of surrounding promoters and enhancers.
Conflict of interest statement
Paul Flicek is a member of the scientific advisory boards of Fabric Genomics, Inc., and Eagle Genomics, Ltd.
Figures




References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

