The Molybdenum Cofactor Biosynthesis Network: In vivo Protein-Protein Interactions of an Actin Associated Multi-Protein Complex
- PMID: 29184564
- PMCID: PMC5694649
- DOI: 10.3389/fpls.2017.01946
The Molybdenum Cofactor Biosynthesis Network: In vivo Protein-Protein Interactions of an Actin Associated Multi-Protein Complex
Abstract
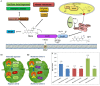
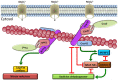
Survival of plants and nearly all organisms depends on the pterin based molybdenum cofactor (Moco) as well as its effective biosynthesis and insertion into apo-enzymes. To this end, both the central Moco biosynthesis enzymes are characterized and the conserved four-step reaction pathway for Moco biosynthesis is well-understood. However, protection mechanisms to prevent degradation during biosynthesis as well as transfer of the highly oxygen sensitive Moco and its intermediates are not fully enlightened. The formation of protein complexes involving transient protein-protein interactions is an efficient strategy for protected metabolic channelling of sensitive molecules. In this review, Moco biosynthesis and allocation network is presented and discussed. This network was intensively studied based on two in vivo interaction methods: bimolecular fluorescence complementation (BiFC) and split-luciferase. Whereas BiFC allows localisation of interacting partners, split-luciferase assay determines interaction strengths in vivo. Results demonstrate (i) interaction of Cnx2 and Cnx3 within the mitochondria and (ii) assembly of a biosynthesis complex including the cytosolic enzymes Cnx5, Cnx6, Cnx7, and Cnx1, which enables a protected transfer of intermediates. The whole complex is associated with actin filaments via Cnx1 as anchor protein. After biosynthesis, Moco needs to be handed over to the specific apo-enzymes. A potential pathway was discovered. Molybdenum-containing enzymes of the sulphite oxidase family interact directly with Cnx1. In contrast, the xanthine oxidoreductase family acquires Moco indirectly via a Moco binding protein (MoBP2) and Moco sulphurase ABA3. In summary, the uncovered interaction matrix enables an efficient transfer for intermediate and product protection via micro-compartmentation.
Keywords: bimolecular fluorescent complementation (BiFC); cytoskeleton; metabolic channelling; molybdenum cofactor; protein-protein interaction network; split-luciferase.
Figures
References
-
- Baillie C. K., Kaufholdt D., Karpinski L. H., Schreiber V., Hänsch S., Evers C., et al. (2016). Detoxification of volcanic sulfur surplus in planta: three different strategies of survival. Environ. Exp. Bot. 126, 44–54. 10.1016/j.envexpbot.2016.02.007 - DOI
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

