A Multilayered Control of the Human Survival Motor Neuron Gene Expression by Alu Elements
- PMID: 29187847
- PMCID: PMC5694776
- DOI: 10.3389/fmicb.2017.02252
A Multilayered Control of the Human Survival Motor Neuron Gene Expression by Alu Elements
Abstract
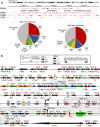
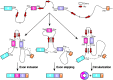
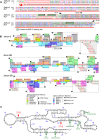
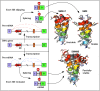
Humans carry two nearly identical copies of Survival Motor Neuron gene: SMN1 and SMN2. Mutations or deletions of SMN1, which codes for SMN, cause spinal muscular atrophy (SMA), a leading genetic disease associated with infant mortality. Aberrant expression or localization of SMN has been also implicated in other pathological conditions, including male infertility, inclusion body myositis, amyotrophic lateral sclerosis and osteoarthritis. SMN2 fails to compensate for the loss of SMN1 due to skipping of exon 7, leading to the production of SMNΔ7, an unstable protein. In addition, SMNΔ7 is less functional due to the lack of a critical C-terminus of the full-length SMN, a multifunctional protein. Alu elements are specific to primates and are generally found within protein coding genes. About 41% of the human SMN gene including promoter region is occupied by more than 60 Alu-like sequences. Here we discuss how such an abundance of Alu-like sequences may contribute toward SMA pathogenesis. We describe the likely impact of Alu elements on expression of SMN. We have recently identified a novel exon 6B, created by exonization of an Alu-element located within SMN intron 6. Irrespective of the exon 7 inclusion or skipping, transcripts harboring exon 6B code for the same SMN6B protein that has altered C-terminus compared to the full-length SMN. We have demonstrated that SMN6B is more stable than SMNΔ7 and likely functions similarly to the full-length SMN. We discuss the possible mechanism(s) of regulation of SMN exon 6B splicing and potential consequences of the generation of exon 6B-containing transcripts.
Keywords: Alu; SMA; SMN; SMN6B; exonization; spinal muscular atrophy; survival motor neuron; transposable elements.
Figures

Similar articles
-
A novel human-specific splice isoform alters the critical C-terminus of Survival Motor Neuron protein.Sci Rep. 2016 Aug 2;6:30778. doi: 10.1038/srep30778. Sci Rep. 2016. PMID: 27481219 Free PMC article.
-
Structural Context of a Critical Exon of Spinal Muscular Atrophy Gene.Front Mol Biosci. 2022 Jul 1;9:928581. doi: 10.3389/fmolb.2022.928581. eCollection 2022. Front Mol Biosci. 2022. PMID: 35847983 Free PMC article. Review.
-
Evolving concepts on human SMN pre-mRNA splicing.RNA Biol. 2007 Jan-Mar;4(1):7-10. doi: 10.4161/rna.4.1.4535. Epub 2007 Jun 4. RNA Biol. 2007. PMID: 17592254
-
Securinine enhances SMN2 exon 7 inclusion in spinal muscular atrophy cells.Biomed Pharmacother. 2017 Apr;88:708-714. doi: 10.1016/j.biopha.2017.01.104. Epub 2017 Jan 30. Biomed Pharmacother. 2017. PMID: 28152480
-
Mechanism of Splicing Regulation of Spinal Muscular Atrophy Genes.Adv Neurobiol. 2018;20:31-61. doi: 10.1007/978-3-319-89689-2_2. Adv Neurobiol. 2018. PMID: 29916015 Free PMC article. Review.
Cited by
-
A survey of transcripts generated by spinal muscular atrophy genes.Biochim Biophys Acta Gene Regul Mech. 2020 Aug;1863(8):194562. doi: 10.1016/j.bbagrm.2020.194562. Epub 2020 May 6. Biochim Biophys Acta Gene Regul Mech. 2020. PMID: 32387331 Free PMC article. Review.
-
ZPR1 prevents R-loop accumulation, upregulates SMN2 expression and rescues spinal muscular atrophy.Brain. 2020 Jan 1;143(1):69-93. doi: 10.1093/brain/awz373. Brain. 2020. PMID: 31828288 Free PMC article.
-
The First Orally Deliverable Small Molecule for the Treatment of Spinal Muscular Atrophy.Neurosci Insights. 2020 Nov 23;15:2633105520973985. doi: 10.1177/2633105520973985. eCollection 2020. Neurosci Insights. 2020. PMID: 33283185 Free PMC article. Review.
-
Transcriptome- and proteome-wide effects of a circular RNA encompassing four early exons of the spinal muscular atrophy genes.Res Sq [Preprint]. 2024 Feb 28:rs.3.rs-3818622. doi: 10.21203/rs.3.rs-3818622/v1. Res Sq. 2024. Update in: Sci Rep. 2024 May 7;14(1):10442. doi: 10.1038/s41598-024-60593-7. PMID: 38464174 Free PMC article. Updated. Preprint.
-
Internal Introns Promote Backsplicing to Generate Circular RNAs from Spinal Muscular Atrophy Gene.Genes (Basel). 2022 Jun 25;13(7):1145. doi: 10.3390/genes13071145. Genes (Basel). 2022. PMID: 35885927 Free PMC article.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

