Structural studies of Chikungunya virus maturation
- PMID: 29203665
- PMCID: PMC5748190
- DOI: 10.1073/pnas.1713166114
Structural studies of Chikungunya virus maturation
Abstract
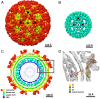
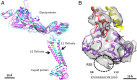
Cleavage of the alphavirus precursor glycoprotein p62 into the E2 and E3 glycoproteins before assembly with the nucleocapsid is the key to producing fusion-competent mature spikes on alphaviruses. Here we present a cryo-EM, 6.8-Å resolution structure of an "immature" Chikungunya virus in which the cleavage site has been mutated to inhibit proteolysis. The spikes in the immature virus have a larger radius and are less compact than in the mature virus. Furthermore, domains B on the E2 glycoproteins have less freedom of movement in the immature virus, keeping the fusion loops protected under domain B. In addition, the nucleocapsid of the immature virus is more compact than in the mature virus, protecting a conserved ribosome-binding site in the capsid protein from exposure. These differences suggest that the posttranslational processing of the spikes and nucleocapsid is necessary to produce infectious virus.
Keywords: Chikungunya virus; alphavirus; conformational changes; cryo-electron microscopy; maturation.
Conflict of interest statement
Conflict of interest statement: M.L.Y., T.K., S.S.H., and M.G.R. declare no competing financial interests. A.U. is an employee of VLP Therapeutics, and W.A. is an officer and shareholder of VLP Therapeutics.
Figures


References
-
- Robinson MC. An epidemic of virus disease in Southern Province, Tanganyika Territory, in 1952-1953. Trans R Soc Trop Med Hyg. 1955;49:28–32. - PubMed
-
- Centers for Disease Control and Prevention 2017 NOWCAST: Chikungunya in the Americas. Available at https://www.cdc.gov/chikungunya/modeling/index.html. Accessed April 11, 2017.
-
- Ryman KD, Klimstra WB. Host responses to alphavirus infection. Immunol Rev. 2008;225:27–45. - PubMed
-
- Kuhn RJ. Togaviridae: The viruses and their replication. In: Knipe DM, Howley PM, editors. Fields Virology. 5th Ed. Lippincott Williams & Wilkins; Philadelphia: 2007. pp. 1001–1022.
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

