A microsatellite diversity analysis and the development of core-set germplasm in a large hulless barley (Hordeum vulgare L.) collection
- PMID: 29207956
- PMCID: PMC5717800
- DOI: 10.1186/s12863-017-0563-x
A microsatellite diversity analysis and the development of core-set germplasm in a large hulless barley (Hordeum vulgare L.) collection
Abstract
Background: Clarifying genetic diversity in a large germplasm resource plays important roles in experimental designs that provides flexible utility in fundamental research and breeding in crops. However, the work is limited due to small collections of barley that are insufficient representatives.
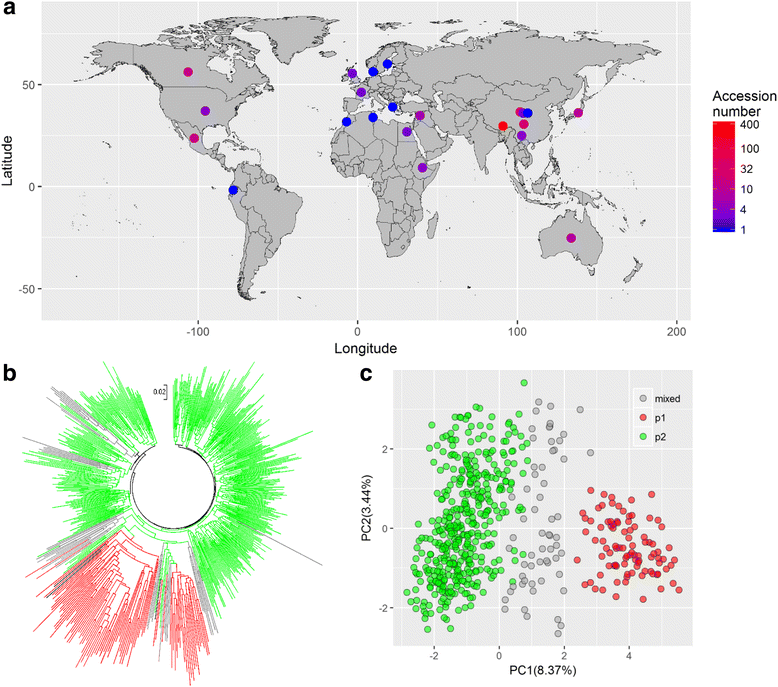
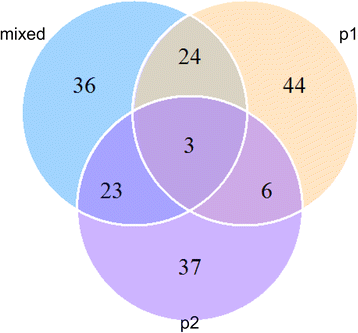
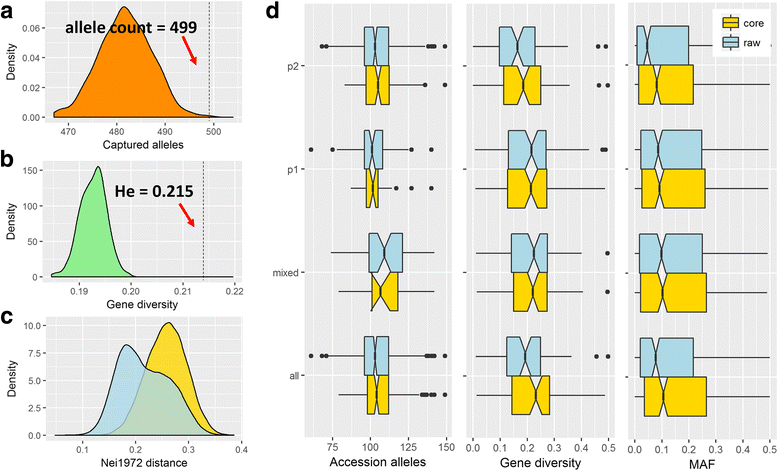
Results: In the present study, we collected 562 hulless barley (Hordeum vulgare L.) accessions with worldwide geographic origins and evaluated their genetic variability and relatedness based on 93 simple sequence repeat (SSR) markers. In an integrated analysis of the population structure, analysis of molecular variance (AMOVA) and pairwise F ST, the 562 barley accessions exhibited a strong stratification that allowed for them to be divided into two major subpopulations (p1 and p2) and an admixture subpopulation, with 93, 408 and 61 accessions, respectively. In a neutral test, considerable proportions of SSR alleles expressed the strong non-neutrality in specific subpopulations (44 and 37), which are probably responsible for population differentiation. To reduce the diversity redundancy in large barley collections, we delicately selected a core set of 200 barley accessions as a tradeoff between diversity and representativeness in an easily handled population. In comparing the 562 barley accessions, the core barley set accounted for 96.2% of allelic diversity and 93% to 95% of phenotypic variability, whereas it exhibited a significant enhancement in minor allelic frequencies, which probably benefit association mapping in the barley core set.
Conclusions: The results provided additional insight into the genetic structure in a large barley germplasm resource, from which an easily manageable barley core set was identified, demonstrating the great potential for discovering key QTLs and ultimately facilitating barley breeding progress.
Keywords: Association mapping; Core germplasm; Genetic diversity; Hulless barley; Population structure.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
References
-
- Takahashi R. The origin and evolution of cultivated barley. Adv Genet. 1955;7:227–266.
-
- Sharma KP, Dahal KR, Basta BK. Proceedings of the 2nd national conference on science and technology, 1994, Kathmandu. Kathmandu: Royal Nepal Acad Sci Tech (RONAST); 1994. Genetic diversity of Nepalese naked barley and possibility of yield improvement; pp. 231–237.
-
- Baniya BK, Dongol DMS, Riley KW. Characterization of Nepalese barley germplasm. Rachis. 1997;16:16–19.
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

