On causal roles and selected effects: our genome is mostly junk
- PMID: 29207982
- PMCID: PMC5718017
- DOI: 10.1186/s12915-017-0460-9
On causal roles and selected effects: our genome is mostly junk
Abstract
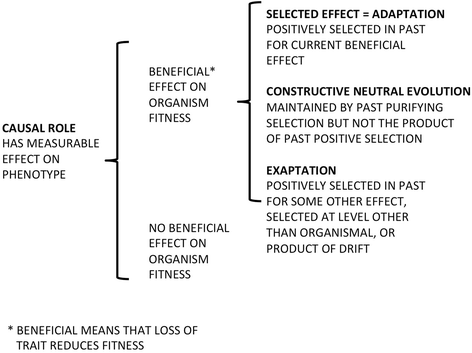
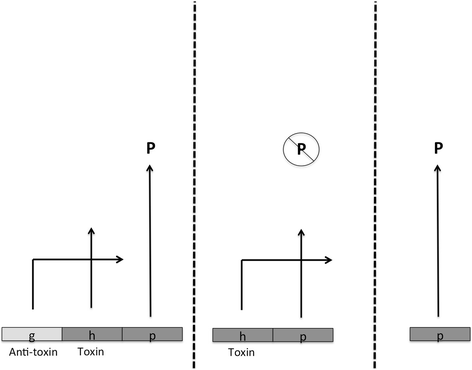
The idea that much of our genome is irrelevant to fitness-is not the product of positive natural selection at the organismal level-remains viable. Claims to the contrary, and specifically that the notion of "junk DNA" should be abandoned, are based on conflating meanings of the word "function". Recent estimates suggest that perhaps 90% of our DNA, though biochemically active, does not contribute to fitness in any sequence-dependent way, and possibly in no way at all. Comparisons to vertebrates with much larger and smaller genomes (the lungfish and the pufferfish) strongly align with such a conclusion, as they have done for the last half-century.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
Can ENCODE tell us how much junk DNA we carry in our genome?Biochem Biophys Res Commun. 2013 Jan 25;430(4):1340-3. doi: 10.1016/j.bbrc.2012.12.074. Epub 2012 Dec 22. Biochem Biophys Res Commun. 2013. PMID: 23268340 Review.
-
Tuning in to the signals: noncoding sequence conservation in vertebrate genomes.Trends Genet. 2008 Jul;24(7):344-52. doi: 10.1016/j.tig.2008.04.005. Epub 2008 May 29. Trends Genet. 2008. PMID: 18514361 Review.
-
Pan-vertebrate conserved non-coding sequences associated with developmental regulation.Brief Funct Genomic Proteomic. 2009 Jul;8(4):256-65. doi: 10.1093/bfgp/elp033. Brief Funct Genomic Proteomic. 2009. PMID: 19752044 Review.
-
Is junk DNA bunk? A critique of ENCODE.Proc Natl Acad Sci U S A. 2013 Apr 2;110(14):5294-300. doi: 10.1073/pnas.1221376110. Epub 2013 Mar 11. Proc Natl Acad Sci U S A. 2013. PMID: 23479647 Free PMC article.
-
Genomic regulatory blocks in vertebrates and implications in human disease.Brief Funct Genomic Proteomic. 2009 Jul;8(4):333-42. doi: 10.1093/bfgp/elp019. Epub 2009 Jun 26. Brief Funct Genomic Proteomic. 2009. PMID: 19561171 Review.
Cited by
-
Epigenetic silencing and genome dynamics determine the fate of giant virus endogenizations in Acanthamoeba.BMC Biol. 2025 Jul 1;23(1):171. doi: 10.1186/s12915-025-02280-1. BMC Biol. 2025. PMID: 40597104 Free PMC article.
-
An Evaluation of Function of Multicopy Noncoding RNAs in Mammals Using ENCODE/FANTOM Data and Comparative Genomics.Mol Biol Evol. 2018 Jun 1;35(6):1451-1462. doi: 10.1093/molbev/msy046. Mol Biol Evol. 2018. PMID: 29617896 Free PMC article.
-
Small Open Reading Frames: How Important Are They for Molecular Evolution?Front Genet. 2020 Oct 20;11:574737. doi: 10.3389/fgene.2020.574737. eCollection 2020. Front Genet. 2020. PMID: 33193682 Free PMC article. No abstract available.
-
DNA sequence-dependent chromatin architecture and nuclear hubs formation.Sci Rep. 2019 Oct 10;9(1):14646. doi: 10.1038/s41598-019-51036-9. Sci Rep. 2019. PMID: 31601866 Free PMC article.
-
Genes and genomes and unnecessary complexity in precision medicine.NPJ Genom Med. 2020 May 4;5:21. doi: 10.1038/s41525-020-0128-1. eCollection 2020. NPJ Genom Med. 2020. PMID: 32377378 Free PMC article. Review.
References
-
- Ohno S. So much “junk” DNA in our genome. In: Smith HH, editor. Evolution of genetic systems. New York: Gordon and Breach; 1972. pp. 366–70.
-
- Lu Q, Bourrat P. The evolutionary gene and the extended evolutionary synthesis. Brit J Phil Sci. 2017. doi.org/10.1093/bjps/axw035.
-
- Cavalier-Smith T. Nuclear volume control by nucleoskeletal DNA, selection for cell volume and cell growth rate, and the solution of the DNA C-value paradox. J Cell Sci. 1978;34:247–78. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

