X-ray diffraction reveals the intrinsic difference in the physical properties of membrane and soluble proteins
- PMID: 29208950
- PMCID: PMC5717104
- DOI: 10.1038/s41598-017-17216-1
X-ray diffraction reveals the intrinsic difference in the physical properties of membrane and soluble proteins
Abstract
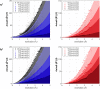
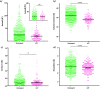
Membrane proteins are distinguished from soluble proteins by their insertion into biological membranes. This insertion is achieved via a noticeable arrangement of hydrophobic amino acids that are exposed at the surface of the protein, and renders the interaction with the aliphatic tails of lipids more energetically favorable. This important difference between these two categories of proteins is the source of the need for a specific handling of membrane proteins, which transpired in the creation of new tools for their recombinant expression, purification and even crystallization. Following this line, we show here that crystals of membrane proteins display systematically higher diffraction anisotropy than those of soluble proteins. This phenomenon dramatically hampers structure solution and refinement, and has a strong impact on the quality of electron-density maps. A farther search for origins of this phenomenon showed that the type of crystallization, and thus the crystal packing, has no impact on anisotropy, nor does the nature or function of the membrane protein. Membrane proteins fully embedded within the membrane display equal anisotropy compared to the ones with extra membranous domains or fusions with soluble proteins. Overall, these results overturn common beliefs and call for a specific handling of their diffraction data.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

