Allosteric pyruvate kinase-based "logic gate" synergistically senses energy and sugar levels in Mycobacterium tuberculosis
- PMID: 29215013
- PMCID: PMC5719368
- DOI: 10.1038/s41467-017-02086-y
Allosteric pyruvate kinase-based "logic gate" synergistically senses energy and sugar levels in Mycobacterium tuberculosis
Abstract
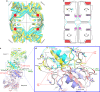
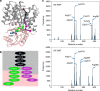
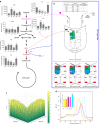
Pyruvate kinase (PYK) is an essential glycolytic enzyme that controls glycolytic flux and is critical for ATP production in all organisms, with tight regulation by multiple metabolites. Yet the allosteric mechanisms governing PYK activity in bacterial pathogens are poorly understood. Here we report biochemical, structural and metabolomic evidence that Mycobacterium tuberculosis (Mtb) PYK uses AMP and glucose-6-phosphate (G6P) as synergistic allosteric activators that function as a molecular "OR logic gate" to tightly regulate energy and glucose metabolism. G6P was found to bind to a previously unknown site adjacent to the canonical site for AMP. Kinetic data and structural network analysis further show that AMP and G6P work synergistically as allosteric activators. Importantly, metabolome profiling in the Mtb surrogate, Mycobacterium bovis BCG, reveals significant changes in AMP and G6P levels during nutrient deprivation, which provides insights into how a PYK OR gate would function during the stress of Mtb infection.
Conflict of interest statement
The authors declare no competing financial interests.
Figures



Similar articles
-
An overview of structure, function, and regulation of pyruvate kinases.Protein Sci. 2019 Oct;28(10):1771-1784. doi: 10.1002/pro.3691. Epub 2019 Aug 12. Protein Sci. 2019. PMID: 31342570 Free PMC article. Review.
-
Pyruvate Kinase Regulates the Pentose-Phosphate Pathway in Response to Hypoxia in Mycobacterium tuberculosis.J Mol Biol. 2019 Sep 6;431(19):3690-3705. doi: 10.1016/j.jmb.2019.07.033. Epub 2019 Aug 2. J Mol Biol. 2019. PMID: 31381898
-
Central Role of Pyruvate Kinase in Carbon Co-catabolism of Mycobacterium tuberculosis.J Biol Chem. 2016 Mar 25;291(13):7060-9. doi: 10.1074/jbc.M115.707430. Epub 2016 Feb 8. J Biol Chem. 2016. PMID: 26858255 Free PMC article.
-
Allosteric regulation of pyruvate kinase from Mycobacterium tuberculosis by metabolites.Biochim Biophys Acta Proteins Proteom. 2019 Feb;1867(2):125-139. doi: 10.1016/j.bbapap.2018.11.002. Epub 2018 Nov 10. Biochim Biophys Acta Proteins Proteom. 2019. PMID: 30419357
-
Expression, purification, and characterization of pyruvate kinase from Mycobacterium tuberculosis: A key allosteric regulatory enzyme.Int J Mycobacteriol. 2018 Oct-Dec;7(4):368-374. doi: 10.4103/ijmy.ijmy_116_18. Int J Mycobacteriol. 2018. PMID: 30531037
Cited by
-
Catalyst-based biomolecular logic gates.Catalysts. 2022 Jul;12(7):712. doi: 10.3390/catal12070712. Epub 2022 Jun 29. Catalysts. 2022. PMID: 37377541 Free PMC article.
-
An overview of structure, function, and regulation of pyruvate kinases.Protein Sci. 2019 Oct;28(10):1771-1784. doi: 10.1002/pro.3691. Epub 2019 Aug 12. Protein Sci. 2019. PMID: 31342570 Free PMC article. Review.
-
PYK-SubstitutionOME: an integrated database containing allosteric coupling, ligand affinity and mutational, structural, pathological, bioinformatic and computational information about pyruvate kinase isozymes.Database (Oxford). 2023 May 3;2023:baad030. doi: 10.1093/database/baad030. Database (Oxford). 2023. PMID: 37171062 Free PMC article.
-
All DACs in a Row: Domain Architectures of Bacterial and Archaeal Diadenylate Cyclases.J Bacteriol. 2023 Apr 25;205(4):e0002323. doi: 10.1128/jb.00023-23. Epub 2023 Apr 6. J Bacteriol. 2023. PMID: 37022175 Free PMC article. Review.
-
Functional cross-talk between allosteric effects of activating and inhibiting ligands underlies PKM2 regulation.Elife. 2019 Jul 2;8:e45068. doi: 10.7554/eLife.45068. Elife. 2019. PMID: 31264961 Free PMC article.
References
-
- World Health Organization. Global Tuberculosis Report 2015. http://www.who.int/tb/publications/global_report/gtbr15_main_text.pdf (2015).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

