FoxO3 an important player in fibrogenesis and therapeutic target for idiopathic pulmonary fibrosis
- PMID: 29217661
- PMCID: PMC5801513
- DOI: 10.15252/emmm.201606261
FoxO3 an important player in fibrogenesis and therapeutic target for idiopathic pulmonary fibrosis
Abstract
Idiopathic pulmonary fibrosis (IPF) is a progressive and fatal parenchymal lung disease with limited therapeutic options, with fibroblast-to-myofibroblast transdifferentiation and hyperproliferation playing a major role. Investigating ex vivo-cultured (myo)fibroblasts from human IPF lungs as well as fibroblasts isolated from bleomycin-challenged mice, Forkhead box O3 (FoxO3) transcription factor was found to be less expressed, hyperphosphorylated, and nuclear-excluded relative to non-diseased controls. Downregulation and/or hyperphosphorylation of FoxO3 was reproduced by exposure of normal human lung fibroblasts to various pro-fibrotic growth factors and cytokines (FCS, PDGF, IGF1, TGF-β1). Moreover, selective knockdown of FoxO3 in the normal human lung fibroblasts reproduced the transdifferentiation and hyperproliferation phenotype. Importantly, mice with global- (Foxo3-/-) or fibroblast-specific (Foxo3f.b-/-) FoxO3 knockout displayed enhanced susceptibility to bleomycin challenge, with augmented fibrosis, loss of lung function, and increased mortality. Activation of FoxO3 with UCN-01, a staurosporine derivative currently investigated in clinical cancer trials, reverted the IPF myofibroblast phenotype in vitro and blocked the bleomycin-induced lung fibrosis in vivo These studies implicate FoxO3 as a critical integrator of pro-fibrotic signaling in lung fibrosis and pharmacological reconstitution of FoxO3 as a novel treatment strategy.
Keywords: fibroblast; forkhead box O transcription factors; idiopathic pulmonary fibrosis; myofibroblast; transdifferentiation.
© 2017 The Authors. Published under the terms of the CC BY 4.0 license.
Figures
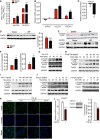
- A
Proliferation rate of N‐HLF (isolated from healthy donor; n = 3) and IPF‐HLF (isolated from IPF patients; n = 3) in the absence or presence of 5% FCS was measured by BrdU incorporation.
- B
mRNA expression of myofibroblasts markers (ACTA2, COL1A1, and COL3A1) in N‐HLF (n = 9) and IPF‐HLF (n = 7) by qPCR.
- C–F
Expression and activity of FoxO3 in N‐HLF and IPF‐HLF (passages 4–5). (C) mRNA expression analysis of FOXO3 by qPCR (n = 7/group). (D, left panel) Representative Western blot of FoxO3. (D, right panel) Densitometrically quantified data of FoxO3/GAPDH expression ratio (n = 6–7/group). (E) Representative Western blots of p‐FoxO3 (Thr32), p‐FoxO3 (Ser253). (F) Densitometrically quantified data of p‐FoxO3 (Thr32) or p‐FoxO3 (Ser253) to GAPDH expression ratio (n = 6–7/group).
- G–J
Representative Western blots of p‐FoxO3 (Thr32), p‐FoxO3 (Ser253), and FoxO3 in serum‐starved (48 h) N‐HLF (n = 3) that were stimulated with 5% FCS (G), PDGF‐BB (H), IGF‐1 (I), or TGF‐β1 (J) as indicated.
- K
Representative immunocytochemistry (ICC) images of FoxO3 localization in N‐HLF (n = 4) stimulated without/with TGF‐β1 (10 ng/ml) as indicated. Scale bar = 50 μm.
- L
mRNA expression of FOXO3 by qPCR (n = 7; left panel) and (middle panel) representative Western blot of FoxO3 in N‐HLF stimulated with TGF‐β1 for 24 h (right panel). Densitometrically quantified data, represented as fold change of FoxO3/GAPDH expression ratio (n = 4).
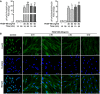
Densitometry quantitation ratio of p‐FoxO3 (Thr32) (left panel) and p‐FoxO3 (Ser253) (right panel) in serum‐starved (48 h) N‐HLF (n = 3) that were stimulated without/with PDGF‐BB as indicated. Quantification is represented as a fold change to control (time corresponded non‐stimulated cells). Data were analyzed using repeated‐measures ANOVA, **P < 0.01 versus control 15 min, § P < 0.05, §§ P < 0.01, §§§ P < 0.001 versus control 30 min.
ICC of FoxO3 in N‐HLF that was serum‐starved for 48 h, and stimulated with PDGF‐BB (60 ng/ml) as indicated. Control image panel represents cells that were left non‐stimulated for 6 h. TO‐PRO3 (blue) was used to label nuclei. FoxO3 and TO‐PRO3 images were overlaid to visualize nuclear and cytoplasmic localization of FoxO3. Images are representative of n = 3. Scale bar = 50 μm.
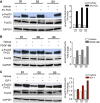
- A–C
Western blots of p‐FoxO3 (Thr32), FoxO3, and GAPDH in serum‐starved (48 h) N‐HLF (n = 3) that was stimulated with 5% FCS (A), PDGF‐BB (B), or IGF‐1 (C) as indicated. Densitometry quantified data of p‐FoxO3 (Thr32) to FoxO3 expression ratios, represented as a fold change to non‐stimulated cells.
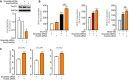
- A
Representative Western blots of FoxO3 in control human lung fibroblasts untransfected or transfected with scramble siRNA or FoxO3 siRNA. Densitometry quantified data of FoxO3/GAPDH expression ratio (n = 3).
- B, C
Human lung fibroblasts from donors (n = 2–3) were transfected with scramble siRNA or FoxO3 siRNA. (B) Transfected cells were stimulated with 5% FCS, IGF‐I (200 ng/ml), and PDGF‐BB (60 ng/ml) or left non‐stimulated, and cell proliferation was measured by BrdU incorporation. (C) mRNA expression of fibroblasts to myofibroblasts markers (ACTA2, COL1A1, and COL3A1) was analyzed by qPCR.

- A, B
Fibroblasts were isolated from saline‐treated mice lungs at day 21 post‐instillation and mice lungs after 14 or 21 days of bleomycin instillation. (A) mRNA expression analysis of Foxo3 by qPCR (n = 4–5/group). (B, left panel) Representative Western blots of p‐FoxO3 (Thr32), FoxO3, and α‐Sma protein levels. Actb was used as a loading control. (B, right panel) Densitometry quantified data of FoxO3, p‐FoxO3 Thr32, and α‐Sma‐to‐Actb expression ratio (n = 4/group).
- C
Percentage of survival of mice challenged with bleomycin. WT (littermates) bleomycin (n = 6 mice), Foxo3 f.b −/− (n = 6 mice) and Foxo3 −/− bleomycin (n = 9 mice).
- D–F
Lung function measurements of mice at day 14 after instillation, (D) total lung capacity, (E) lung compliance, and (F) lung tissue resistance.
- G
Hydroxyproline levels in mouse lungs.
- H
Representative H&E staining of whole left lung (upper panel) and higher magnification (lower panel). Scale bar = 100 μm.
- I
Fibrotic score.

- A–D
Immunofluorescence staining was performed on WT, Foxo3 −/− and Foxo3 f.b −/− mice lung sections (saline‐ and bleomycin‐instilled) using CD68, CD45, and CD3 antibodies. Representative pictographs depicting CD68 (A), CD45 (B), and CD3 (C) staining in green from mice (n = 3) in each group. DAPI was used as a nuclear stain. Scale bar = 50 μm. Fluorescence intensities of CD3‐stained sections (n = 5/6 per group) were quantified using ImageJ software and normalized to DAPI intensity. Data are expressed as mean ± SEM and were analyzed using repeated‐measures one‐way ANOVA, ***P < 0.001 versus WT saline group and §§ P < 0.01 versus WT bleomycin group. m1, m2, and m3 represent three different mice evaluated in each group.
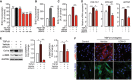
- A–C
Serum‐starved (48 h) N‐HLF (n = 3) were stimulated with 5% FCS (A), PDGF‐BB (60 ng/ml) (B), or IGF‐1 (200 ng/ml) (C) in the presence of UCN‐01 or vehicle (DMSO) as indicated and cell proliferation was measured by BrdU incorporation after 24 h. Data represent percentage of control; vehicle‐treated cells (A) or serum‐starved cells (B and C).
- D–F
Serum‐starved (48 h) N‐HLF (n = 3) were left non‐stimulated or stimulated with TGF‐β1 (10 ng/ml) in medium containing UCN‐01 (50 nM) or vehicle (DMSO) for an additional 24 h and the following analyses were performed: (D) mRNA expression analysis of fibroblast‐to‐myofibroblasts markers (COL1A1, COL3A1, and ACTA2) expression by qPCR. (E, left panel) Representative Western blots of collagen 1a (Col1a) and α‐SMA. GAPDH was used as a loading control. (E, right panel) Densitometry quantified data of Col1a and α‐SMA‐to‐GAPDH expression ratio. (F) ICC of α‐SMA (red, upper) and Col1a (green, lower). TO‐PRO3 (blue) was used to label nuclei. Scale bar = 50 μm.

- A–I
Serum‐starved (48 h) N‐HLF were stimulated with 5% FCS (A–C), IGF‐1 (200 ng/ml) (D–F), or PDGF‐BB (60 ng/ml) (G–I) in medium containing UCN‐01 (50 nM) or wortmannin (Wort.) (500 nM) or vehicle (DMSO) for 30 min and Western blot analysis was performed with the antibodies as described. (A, D, and G) Representative Western blots of p‐FoxO3 (Thr32), p‐FoxO3 (Ser253), FoxO3, p‐AKT (Thr308), AKT, and GAPDH. (B, C, E, F, H, and I) Densitometry quantified data of p‐FoxO3 (Thr32) or p‐FoxO3 (Ser253) to FoxO3 and p‐AKT (Thr308) to AKT expression ratios, represented as a fold change to non‐stimulated cells.
- J
ICC of FoxO3 in N‐HLF that were serum‐starved for 48 h and were stimulated with 5% FCS or PDGF‐BB (60 ng/ml) or IGF‐1 (200 ng/ml) and treated with 50 nM UCN‐01 or 500 nM wortmannin or vehicle control (DMSO) for 6 h. TO‐PRO3 (blue) was used to label nuclei. FoxO3 and TO‐PRO3 images were overlaid to visualize nuclear and cytoplasmic localization of FoxO3. Images are representative of n = 3. Scale bar = 50 μm.

- A–D
Forty‐eight hours serum‐starved N‐HLF were left non‐stimulated or stimulated with TGF‐β1 (10 ng/ml) in medium containing UCN‐01 (50 nM) or vehicle (DMSO) for 24 h. (A) mRNA expression of FOXO3 by qPCR. Data are expressed as mean ± SEM and were analyzed using one‐way ANOVA, **P < 0.01, ***P < 0.001 versus vehicle‐treated cells. (B) Representative Western blots of p‐FoxO3 (Thr32), p‐FoxO3 (Ser253), FoxO3, p‐AKT (Thr308), AKT and GAPDH. (C, D) Densitometry quantified data, represented as fold change (n = 3). In all panels, cells were serum‐starved for 48 h before stimulations. Data are expressed as mean ± SEM and were analyzed using repeated‐measures one‐way ANOVA, n.s. = not significant, *P < 0.05, **P < 0.01, ***P < 0.001 versus vehicle‐treated cells.
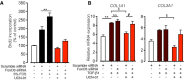
N‐HLF were transfected with scramble siRNA or FoxO3 siRNA. 6 h after transfection, cells were serum‐starved for 36 h and then stimulated with 5% FCS in the presence or absence of UCN‐01 (50 nM), and cell proliferation was measured by BrdU incorporation after 24 h. Scramble siRNA‐transfected cells were left non‐stimulated for 24 h as an additional control. Data represent percentage of control, scramble siRNA non‐stimulated cells (n = 2–3). Data are expressed as mean ± SEM and were analyzed using one‐way ANOVA, **P < 0.01 versus 5% FCS‐scramble siRNA.
N‐HLF was transfected with scramble siRNA or FoxO3 siRNA. 6 h after transfection, cells were serum‐starved for 36 h and then stimulated with TGF‐β1 (10 ng/ml) in the presence or absence of UCN‐01 (50 nM) for 24 h and mRNA expression of COL1A1 and COL3A1 were measured with qPCR (n = 3). Data are expressed as mean ± SEM and were analyzed using one‐way ANOVA, n.s. = not significant, **P < 0.01 versus scramble siRNA, § P < 0.05 versus TGF‐β1‐scramble siRNA, # P < 0.05 versus UCN‐01‐scramble siRNA.
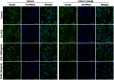
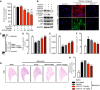
- A
Serum‐starved (48 h) IPF‐HLF (n = 5) were stimulated with 5% FCS and treated with UCN‐01 (10, 50 100, and 200 nM) or vehicle (DMSO) or left untreated, and cell proliferation was measured by BrdU incorporation after 24 h.
- B
IPF‐HLF cells that were serum‐starved for 48 h and stimulated with TGF‐β1 (10 ng/ml) and treated with UCN‐01 (50 nM) or vehicle (DMSO) for 24 h. (B, left panel) Western blots of p‐FoxO3 (Thr32), p‐FoxO3 (Ser253), FoxO3, p‐AKT (Thr308), AKT, and GAPDH. (B, right panel) ICC of α‐SMA (red) and Col1a (green). TO‐PRO3 (blue) was used to label nuclei. Scale bar = 50 μm. Images are representative of n = 3.
- C
Scheme shows experimental setup.
- D–F
Lung function measurements of mice, (D) total lung capacity, (E) lung compliance, and (F) tissue resistance.
- G
Representative H&E staining of whole left lung.
- H
Fibrotic score.

- A–C
Lung function measurements of mice, (A) total lung capacity, (B) lung compliance, and (C) tissue resistance.
- D
Hydroxyproline levels in mice lungs.
- E, F
Stimulation with TGF‐β1 (10 ng/ml) in the presence or absence of UCN‐01 (50 nM) for 24 h, followed by (E) Western blotting for p‐FoxO3 (Thr32), FoxO3, AKT, GAPDH, and (F) qPCRs for COL1A1 and COL3A1 (n = 3).
- G
Model of signaling events involving FoxO3 during IPF pathogenesis. In lung fibroblasts, FoxO3 with nuclear localization leads to transcription of genes involved in regulation of apoptosis, migration, differentiation, and cell‐cycle inhibition. Pro‐fibrotic factors (TGF‐β1, IGF‐1, PDGF‐BB) bind to their receptors and activate PI3K, followed by phosphorylation of AKT. AKT phosphorylates and thereby inactivates FoxO3, followed by nuclear exclusion and degradation of FoxO3, resulting in progression of lung fibrosis. UCN‐01 inhibits AKT phosphorylation resulting in FoxO3 reactivation and attenuation of disease progression. Transforming growth factor β receptor (TGF‐βR), transforming growth factor β1 (TGF‐β1), platelet‐derived growth factor receptor (PDGFR), platelet‐derived growth factor‐BB (PDGF‐BB), insulin‐like growth factor receptor (IGFR), and insulin‐like growth factor‐1 (IGF‐1).
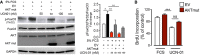
- A, B
N‐HLF was transfected with empty vector (EV) or AKT mutant (AKT mut) plasmid. 6 h after transfection, cells were serum‐starved for 36 h and then stimulated with 5% FCS in the presence or absence of UCN‐01. From the above‐treated samples, after 24 h, Western blots [p‐FoxO3 (Thr32), FoxO3, AKT, AKT mut, GAPDH] and cell proliferation measurements (BrdU incorporation) were performed. Data represent percentage of control, EV non‐stimulated cells (n = 3). Data are expressed as mean ± SEM and were analyzed using one‐way ANOVA, *P < 0.05, **P < 0.01, and ***P < 0.001 versus 5% FCS‐EV, and §§§ P < 0.001 versus UCN‐01‐EV.
References
-
- Adachi M, Osawa Y, Uchinami H, Kitamura T, Accili D, Brenner DA (2007) The forkhead transcription factor FoxO1 regulates proliferation and transdifferentiation of hepatic stellate cells. Gastroenterology 132: 1434–1446 - PubMed
-
- Bonner JC (2004) Regulation of PDGF and its receptors in fibrotic diseases. Cytokine Growth Factor Rev 15: 255–273 - PubMed
-
- Brunet A, Bonni A, Zigmond MJ, Lin MZ, Juo P, Hu LS, Anderson MJ, Arden KC, Blenis J, Greenberg ME (1999) Akt promotes cell survival by phosphorylating and inhibiting a Forkhead transcription factor. Cell 96: 857–868 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

