Identification of UDP-glucose binding site in glycosyltransferase domain of sucrose phosphate synthase from sugarcane (Saccharum officinarum) by structure-based site-directed mutagenesis
- PMID: 29222806
- PMCID: PMC5899720
- DOI: 10.1007/s12551-017-0360-9
Identification of UDP-glucose binding site in glycosyltransferase domain of sucrose phosphate synthase from sugarcane (Saccharum officinarum) by structure-based site-directed mutagenesis
Abstract
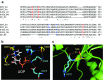
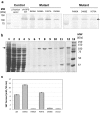
Sucrose phosphate synthase (SPS) is believed to be the key enzyme for controlling the biosynthesis of sucrose. SPSs consist of a functional glycosyltransferase domain that shares conserved residues with the glycosyltransferase domain of sucrose biosynthesis-related protein. The formation of sucrose-6-phosphate is catalyzed by SPS with the transfer of a glycosyl group of uridine diphosphate glucose (UDP-G) as an activated donor sugar to a fructose-6-phosphate as a sugar acceptor. However, understanding of the mechanism of catalytic and substrate binding in SPS is very limited. Based on amino acid sequence alignments with several enzymes that belong to the glycosyltransferase family, the UDP-G binding sites that might be critical for catalytic mechanism were identified. Here, we report that single point mutation of R496, D498, and V570 located in the proposed UDP-G binding site led to less active or complete loss of enzyme activity. Through structure-based site-directed mutagenesis and biochemical studies, the results indicated that these residues contribute to the catalytic activity of plant SPS. Moreover, understanding of the UDP-G binding site provides an insight into new strategies for enzyme engineering and redesigning a catalytic mechanism for UDP.
Keywords: Glycosyltransferase; Site-directed mutagenesis; Sucrose phosphate synthase; Sugarcane; Uridine diphosphate glucose.
Conflict of interest statement
Conflict of interest
Widhi Dyah Sawitri declares that she has no conflicts of interest. Siti Nurul Afidah declares that she has no conflicts of interest. Atsushi Nakagawa declares that he has no conflicts of interest. Toshiharu Hase declares that he has no conflicts of interest. Bambang Sugiharto declares that he has no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Figures
Similar articles
-
Mutation of UDP-glucose binding motif residues lead to increased affinity for ADP-glucose in sugarcane sucrose phosphate synthase.Mol Biol Rep. 2021 Feb;48(2):1697-1706. doi: 10.1007/s11033-021-06181-8. Epub 2021 Feb 2. Mol Biol Rep. 2021. PMID: 33528727
-
The structure of sucrose phosphate synthase from Halothermothrix orenii reveals its mechanism of action and binding mode.Plant Cell. 2008 Apr;20(4):1059-72. doi: 10.1105/tpc.107.051193. Epub 2008 Apr 18. Plant Cell. 2008. PMID: 18424616 Free PMC article.
-
New insight into the catalytic properties of rice sucrose synthase.Plant Mol Biol. 2016 Jan;90(1-2):127-35. doi: 10.1007/s11103-015-0401-3. Epub 2015 Oct 31. Plant Mol Biol. 2016. PMID: 26520834
-
Exploring the active site in UDP-glucose pyrophosphorylase by affinity labelling and site-directed mutagenesis.Biotechnol Appl Biochem. 1993 Oct;18(2):209-16. Biotechnol Appl Biochem. 1993. PMID: 8251118 Review.
-
[Function, structure and catalytic mechanism of sucrose phosphate synthase: a review].Sheng Wu Gong Cheng Xue Bao. 2021 Jun 25;37(6):1858-1868. doi: 10.13345/j.cjb.200743. Sheng Wu Gong Cheng Xue Bao. 2021. PMID: 34227281 Review. Chinese.
Cited by
-
Essential Oil Derived from Underutilized Plants Cymbopogon khasianus Poses Diverse Biological Activities against "Aspergillosis" and "Mucormycosis".Russ Agric Sci. 2023;49(2):172-183. doi: 10.3103/S106836742302012X. Epub 2023 May 17. Russ Agric Sci. 2023. PMID: 37220552 Free PMC article.
-
Essential oil from Cymbopogon citratus exhibits "anti-aspergillosis" potential: in-silico molecular docking and in vitro studies.Bull Natl Res Cent. 2022;46(1):23. doi: 10.1186/s42269-022-00711-5. Epub 2022 Jan 29. Bull Natl Res Cent. 2022. PMID: 35125860 Free PMC article.
-
Current perspectives on the regulatory mechanisms of sucrose accumulation in sugarcane.Heliyon. 2024 Feb 28;10(5):e27277. doi: 10.1016/j.heliyon.2024.e27277. eCollection 2024 Mar 15. Heliyon. 2024. PMID: 38463882 Free PMC article. Review.
-
Comparative Transcriptome Analysis of Gene Expression and Regulatory Characteristics Associated with Different Vernalization Periods in Brassica rapa.Genes (Basel). 2020 Apr 5;11(4):392. doi: 10.3390/genes11040392. Genes (Basel). 2020. PMID: 32260536 Free PMC article.
-
Mutation of UDP-glucose binding motif residues lead to increased affinity for ADP-glucose in sugarcane sucrose phosphate synthase.Mol Biol Rep. 2021 Feb;48(2):1697-1706. doi: 10.1007/s11033-021-06181-8. Epub 2021 Feb 2. Mol Biol Rep. 2021. PMID: 33528727
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

