Successful Establishment of Plasmids R1 and pMV158 in a New Host Requires the Relief of the Transcriptional Repression of Their Essential rep Genes
- PMID: 29250051
- PMCID: PMC5717011
- DOI: 10.3389/fmicb.2017.02367
Successful Establishment of Plasmids R1 and pMV158 in a New Host Requires the Relief of the Transcriptional Repression of Their Essential rep Genes
Abstract
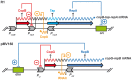
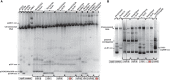
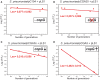
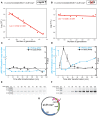
Although differing in size, encoded traits, host range, and replication mechanism, both narrow-host-range theta-type conjugative enterobacterial plasmid R1 and promiscuous rolling-circle-type mobilizable streptococcal plasmid pMV158 encode a transcriptional repressor protein, namely CopB in R1 and CopG in pMV158, involved in replication control. The gene encoding CopB or CopG is cotranscribed with a downstream gene that encodes the replication initiator Rep protein of the corresponding plasmid. However, whereas CopG is an auto-repressor that inhibits transcription of the entire copG-repB operon, CopB is expressed constitutively and represses a second, downstream promoter that directs transcription of repA. As a consequence of the distinct regulatory pathways implied by CopB and CopG, these repressor proteins play a different role in control of plasmid replication during the steady state: while CopB has an auxiliary role by keeping repressed the regulated promoter whenever the plasmid copy number is above a low threshold, CopG plays a primary role by acting coordinately with RNAII. Here, we have studied the role of the regulatory circuit mediated by these transcriptional repressors during the establishment of these two plasmids in a new host cell, and found that excess Cop repressor molecules in the recipient cell result in a severe decrease in the frequency and/or the velocity of appearance of transformant colonies for the cognate plasmid but not for unrelated plasmids. Using the pMV158 replicon as a model system, together with highly sensitive real-time qPCR and inverse PCR methods, we have also analyzed the effect of CopG on the kinetics of repopulation of the plasmid in Streptococcus pneumoniae. We show that, whereas in the absence of CopG pMV158 repopulation occurs mainly during the first 45 min following plasmid transfer, the presence of the transcriptional repressor in the recipient cell severely impairs the replicon repopulation and makes the plasmid replicate at approximately the same rate as the chromosome at any time after transformation, which results in maximal plasmid loss rate in the absence of selection. Overall, these findings indicate that unrepressed activity of the Cop-regulated promoter is crucial for the successful colonization of the recipient bacterial cells by the plasmid.
Keywords: Cop transcriptional repressors; R1 replicon; establishment phase replication; pMV158 replicon; plasmid replication rate; plasmid repopulation.
Figures


Similar articles
-
Translation initiation of the replication initiator repB gene of promiscuous plasmid pMV158 is led by an extended non-SD sequence.Plasmid. 2013 Jul;70(1):69-77. doi: 10.1016/j.plasmid.2013.01.011. Epub 2013 Feb 16. Plasmid. 2013. PMID: 23419647
-
Plasmid transcriptional repressor CopG oligomerises to render helical superstructures unbound and in complexes with oligonucleotides.J Mol Biol. 2001 Jul 6;310(2):403-17. doi: 10.1006/jmbi.2001.4760. J Mol Biol. 2001. PMID: 11428897
-
Structural features of the plasmid pMV158-encoded transcriptional repressor CopG, a protein sharing similarities with both helix-turn-helix and beta-sheet DNA binding proteins.Proteins. 1998 Aug 1;32(2):248-61. doi: 10.1002/(sici)1097-0134(19980801)32:2<248::aid-prot11>3.0.co;2-d. Proteins. 1998. PMID: 9714164
-
Bringing them together: plasmid pMV158 rolling circle replication and conjugation under an evolutionary perspective.Plasmid. 2014 Jul;74:15-31. doi: 10.1016/j.plasmid.2014.05.004. Epub 2014 Jun 2. Plasmid. 2014. PMID: 24942190 Free PMC article. Review.
-
The Facts and Family Secrets of Plasmids That Replicate via the Rolling-Circle Mechanism.Microbiol Mol Biol Rev. 2022 Mar 16;86(1):e0022220. doi: 10.1128/MMBR.00222-20. Epub 2021 Dec 8. Microbiol Mol Biol Rev. 2022. PMID: 34878299 Free PMC article. Review.
Cited by
-
Acidic pH Decreases the Endonuclease Activity of Initiator RepB and Increases the Stability of the Covalent RepB-DNA Intermediate while Has Only a Limited Effect on the Replication of Plasmid pMV158 in Lactococcus lactis.Front Mol Biosci. 2021 Mar 5;8:634461. doi: 10.3389/fmolb.2021.634461. eCollection 2021. Front Mol Biosci. 2021. PMID: 33889596 Free PMC article.
-
Plasmid copy number as a modulator in bacterial pathogenesis and antibiotic resistance.NPJ Antimicrob Resist. 2025 Aug 18;3(1):72. doi: 10.1038/s44259-025-00145-9. NPJ Antimicrob Resist. 2025. PMID: 40826263 Free PMC article. Review.
-
Molecular dissection of the replication system of plasmid pIGRK encoding two in-frame Rep proteins with antagonistic functions.BMC Microbiol. 2019 Nov 13;19(1):254. doi: 10.1186/s12866-019-1595-3. BMC Microbiol. 2019. PMID: 31722681 Free PMC article.
-
Combining Modules for Versatile and Optimal Labeling of Lactic Acid Bacteria: Two pMV158-Family Promiscuous Replicons, a Pneumococcal System for Constitutive or Inducible Gene Expression, and Two Fluorescent Proteins.Front Microbiol. 2019 Jun 26;10:1431. doi: 10.3389/fmicb.2019.01431. eCollection 2019. Front Microbiol. 2019. PMID: 31297101 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

