Mapping genetic variations to three-dimensional protein structures to enhance variant interpretation: a proposed framework
- PMID: 29254494
- PMCID: PMC5735928
- DOI: 10.1186/s13073-017-0509-y
Mapping genetic variations to three-dimensional protein structures to enhance variant interpretation: a proposed framework
Abstract
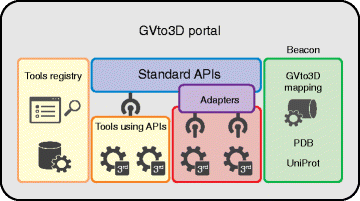
The translation of personal genomics to precision medicine depends on the accurate interpretation of the multitude of genetic variants observed for each individual. However, even when genetic variants are predicted to modify a protein, their functional implications may be unclear. Many diseases are caused by genetic variants affecting important protein features, such as enzyme active sites or interaction interfaces. The scientific community has catalogued millions of genetic variants in genomic databases and thousands of protein structures in the Protein Data Bank. Mapping mutations onto three-dimensional (3D) structures enables atomic-level analyses of protein positions that may be important for the stability or formation of interactions; these may explain the effect of mutations and in some cases even open a path for targeted drug development. To accelerate progress in the integration of these data types, we held a two-day Gene Variation to 3D (GVto3D) workshop to report on the latest advances and to discuss unmet needs. The overarching goal of the workshop was to address the question: what can be done together as a community to advance the integration of genetic variants and 3D protein structures that could not be done by a single investigator or laboratory? Here we describe the workshop outcomes, review the state of the field, and propose the development of a framework with which to promote progress in this arena. The framework will include a set of standard formats, common ontologies, a common application programming interface to enable interoperation of the resources, and a Tool Registry to make it easy to find and apply the tools to specific analysis problems. Interoperability will enable integration of diverse data sources and tools and collaborative development of variant effect prediction methods.
Conflict of interest statement
Competing interests
GG holds stock options in Arivale, Inc. Arivale, Inc. did not fund the study and was not involved in its design, implementation, or reporting. MH is an employee at Human Longevity, Inc. The other authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
References
Publication types
MeSH terms
Grants and funding
- OT3 TR002026/TR/NCATS NIH HHS/United States
- IIS-1636804/National Science Foundation
- OT3TR002026/TR/NCATS NIH HHS/United States
- BB/J019364/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- P30-CA008748/CA/NCI NIH HHS/United States
- P30 CA008748/CA/NCI NIH HHS/United States
- IIS-1636903/National Science Foundation
- U54 HG007990/HG/NHGRI NIH HHS/United States
- BB/G022682/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/L020742/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- U24 CA204817/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources

