Genomic Prediction and Association Mapping of Curd-Related Traits in Gene Bank Accessions of Cauliflower
- PMID: 29255118
- PMCID: PMC5919744
- DOI: 10.1534/g3.117.300199
Genomic Prediction and Association Mapping of Curd-Related Traits in Gene Bank Accessions of Cauliflower
Abstract
Genetic resources are an important source of genetic variation for plant breeding. Genome-wide association studies (GWAS) and genomic prediction greatly facilitate the analysis and utilization of useful genetic diversity for improving complex phenotypic traits in crop plants. We explored the potential of GWAS and genomic prediction for improving curd-related traits in cauliflower (Brassica oleracea var. botrytis) by combining 174 randomly selected cauliflower gene bank accessions from two different gene banks. The collection was genotyped with genotyping-by-sequencing (GBS) and phenotyped for six curd-related traits at two locations and three growing seasons. A GWAS analysis based on 120,693 single-nucleotide polymorphisms identified a total of 24 significant associations for curd-related traits. The potential for genomic prediction was assessed with a genomic best linear unbiased prediction model and BayesB. Prediction abilities ranged from 0.10 to 0.66 for different traits and did not differ between prediction methods. Imputation of missing genotypes only slightly improved prediction ability. Our results demonstrate that GWAS and genomic prediction in combination with GBS and phenotyping of highly heritable traits can be used to identify useful quantitative trait loci and genotypes among genetically diverse gene bank material for subsequent utilization as genetic resources in cauliflower breeding.
Keywords: GenPred; Genomic Selection; Shared Data Resources; cauliflower; gene bank; genome-wide association study; genomic prediction; genotyping-by-sequencing.
Copyright © 2018 Thorwarth et al.
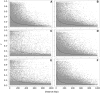
Figures


Similar articles
-
Genotype Imputation in Winter Wheat Using First-Generation Haplotype Map SNPs Improves Genome-Wide Association Mapping and Genomic Prediction of Traits.G3 (Bethesda). 2019 Jan 9;9(1):125-133. doi: 10.1534/g3.118.200664. G3 (Bethesda). 2019. PMID: 30420469 Free PMC article.
-
Genome-wide association study and selective sweep analysis uncover candidate genes controlling curd branch length in cauliflower.Theor Appl Genet. 2024 Aug 28;137(9):209. doi: 10.1007/s00122-024-04719-5. Theor Appl Genet. 2024. PMID: 39196430
-
Understanding population structure and detection of QTLs for curding-related traits in Indian cauliflower by genotyping by sequencing analysis.Funct Integr Genomics. 2021 Nov;21(5-6):679-693. doi: 10.1007/s10142-021-00811-x. Epub 2021 Oct 18. Funct Integr Genomics. 2021. PMID: 34664160
-
Next generation breeding.Plant Sci. 2016 Jan;242:3-13. doi: 10.1016/j.plantsci.2015.07.010. Epub 2015 Jul 19. Plant Sci. 2016. PMID: 26566820 Review.
-
Advances and Challenges for QTL Analysis and GWAS in the Plant-Breeding of High-Yielding: A Focus on Rapeseed.Biomolecules. 2021 Oct 15;11(10):1516. doi: 10.3390/biom11101516. Biomolecules. 2021. PMID: 34680149 Free PMC article. Review.
Cited by
-
Genomic Selection in Sugarcane: Current Status and Future Prospects.Front Plant Sci. 2021 Sep 27;12:708233. doi: 10.3389/fpls.2021.708233. eCollection 2021. Front Plant Sci. 2021. PMID: 34646284 Free PMC article. Review.
-
Genome-Wide Association and Regional Heritability Mapping of Plant Architecture, Lodging and Productivity in Phaseolus vulgaris.G3 (Bethesda). 2018 Jul 31;8(8):2841-2854. doi: 10.1534/g3.118.200493. G3 (Bethesda). 2018. PMID: 29967054 Free PMC article.
-
Using Genome-Wide Predictions to Assess the Phenotypic Variation of a Barley (Hordeum sp.) Gene Bank Collection for Important Agronomic Traits and Passport Information.Front Plant Sci. 2021 Jan 11;11:604781. doi: 10.3389/fpls.2020.604781. eCollection 2020. Front Plant Sci. 2021. PMID: 33505414 Free PMC article.
-
Hybrid Prediction in Horticulture Crop Breeding: Progress and Challenges.Plants (Basel). 2024 Oct 4;13(19):2790. doi: 10.3390/plants13192790. Plants (Basel). 2024. PMID: 39409660 Free PMC article. Review.
-
A Case of Need: Linking Traits to Genebank Accessions.Biopreserv Biobank. 2018 Oct;16(5):337-349. doi: 10.1089/bio.2018.0033. Biopreserv Biobank. 2018. PMID: 30325668 Free PMC article. Review.
References
-
- Bao Y., Vuong T., Meinhardt C., Tiffin P., Denny R., et al. , 2014. Potential of association mapping and genomic selection to explore PI 88788 derived soybean cyst nematode resistance. Plant Genome 7 DOI:10.3835/plantgenome2013.11.0039
-
- Bates D., Maechler M., Bolker B., Walker S., 2014. lme4: linear mixed-effects models using Eigen and S4. R package version 1: 1–23.
-
- Benjamini Y., Hochberg Y., 1995. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57: 289–300.
-
- Bevan M. W., Uauy C., Wulff B. B., Zhou J., Krasileva K., et al. , 2017. Genomic innovation for crop improvement. Nature 543: 346–354. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
