Sexual-lineage-specific DNA methylation regulates meiosis in Arabidopsis
- PMID: 29255257
- PMCID: PMC7611288
- DOI: 10.1038/s41588-017-0008-5
Sexual-lineage-specific DNA methylation regulates meiosis in Arabidopsis
Abstract
DNA methylation regulates eukaryotic gene expression and is extensively reprogrammed during animal development. However, whether developmental methylation reprogramming during the sporophytic life cycle of flowering plants regulates genes is presently unknown. Here we report a distinctive gene-targeted RNA-directed DNA methylation (RdDM) activity in the Arabidopsis thaliana male sexual lineage that regulates gene expression in meiocytes. Loss of sexual-lineage-specific RdDM causes mis-splicing of the MPS1 gene (also known as PRD2), thereby disrupting meiosis. Our results establish a regulatory paradigm in which de novo methylation creates a cell-lineage-specific epigenetic signature that controls gene expression and contributes to cellular function in flowering plants.
Conflict of interest statement
The authors declare no competing financial interests.
Figures


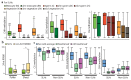




Similar articles
-
Accurate Chromosome Segregation at First Meiotic Division Requires AGO4, a Protein Involved in RNA-Dependent DNA Methylation in Arabidopsis thaliana.Genetics. 2016 Oct;204(2):543-553. doi: 10.1534/genetics.116.189217. Epub 2016 Jul 27. Genetics. 2016. PMID: 27466226 Free PMC article.
-
The cytological and molecular role of domains rearranged methyltransferase3 in RNA-dependent DNA methylation of Arabidopsis thaliana.BMC Res Notes. 2014 Oct 14;7:721. doi: 10.1186/1756-0500-7-721. BMC Res Notes. 2014. PMID: 25316414 Free PMC article.
-
MSI4/FVE is required for accumulation of 24-nt siRNAs and DNA methylation at a subset of target regions of RNA-directed DNA methylation.Plant J. 2021 Oct;108(2):347-357. doi: 10.1111/tpj.15441. Epub 2021 Aug 21. Plant J. 2021. PMID: 34314526 Free PMC article.
-
Mechanisms underlying epigenetic regulation in Arabidopsis thaliana.Integr Comp Biol. 2014 Jul;54(1):61-7. doi: 10.1093/icb/icu030. Epub 2014 May 7. Integr Comp Biol. 2014. PMID: 24808013 Free PMC article. Review.
-
RNA-directed DNA methylation in plants: Where to start?RNA Biol. 2013 Oct;10(10):1593-6. doi: 10.4161/rna.26312. RNA Biol. 2013. PMID: 25003825 Free PMC article. Review.
Cited by
-
Partial maintenance of organ-specific epigenetic marks during plant asexual reproduction leads to heritable phenotypic variation.Proc Natl Acad Sci U S A. 2018 Sep 25;115(39):E9145-E9152. doi: 10.1073/pnas.1805371115. Epub 2018 Sep 10. Proc Natl Acad Sci U S A. 2018. PMID: 30201727 Free PMC article.
-
Divergent DNA Methylation Signatures of Juvenile Seedlings, Grafts and Adult Apple Trees.Epigenomes. 2020 Mar 1;4(1):4. doi: 10.3390/epigenomes4010004. Epigenomes. 2020. PMID: 34968238 Free PMC article.
-
License to Regulate: Noncoding RNA Special Agents in Plant Meiosis and Reproduction.Front Plant Sci. 2021 Aug 20;12:662185. doi: 10.3389/fpls.2021.662185. eCollection 2021. Front Plant Sci. 2021. PMID: 34489987 Free PMC article. Review.
-
Quantitative modelling of fine-scale variations in the Arabidopsis thaliana crossover landscape.Quant Plant Biol. 2022 Feb 15;3:e3. doi: 10.1017/qpb.2021.17. eCollection 2022. Quant Plant Biol. 2022. PMID: 37077963 Free PMC article.
-
DNA methylation dynamics in male germline development in Brassica Rapa.Mol Hortic. 2025 Mar 4;5(1):16. doi: 10.1186/s43897-024-00137-9. Mol Hortic. 2025. PMID: 40033451 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

