Structural basis for substrate specificity of methylsuccinyl-CoA dehydrogenase, an unusual member of the acyl-CoA dehydrogenase family
- PMID: 29275330
- PMCID: PMC5798300
- DOI: 10.1074/jbc.RA117.000764
Structural basis for substrate specificity of methylsuccinyl-CoA dehydrogenase, an unusual member of the acyl-CoA dehydrogenase family
Abstract
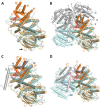
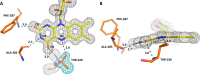
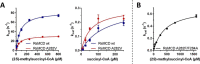
(2S)-methylsuccinyl-CoA dehydrogenase (MCD) belongs to the family of FAD-dependent acyl-CoA dehydrogenase (ACD) and is a key enzyme of the ethylmalonyl-CoA pathway for acetate assimilation. It catalyzes the oxidation of (2S)-methylsuccinyl-CoA to α,β-unsaturated mesaconyl-CoA and shows only about 0.5% activity with succinyl-CoA. Here we report the crystal structure of MCD at a resolution of 1.37 Å. The enzyme forms a homodimer of two 60-kDa subunits. Compared with other ACDs, MCD contains an ∼170-residue-long N-terminal extension that structurally mimics a dimer-dimer interface of these enzymes that are canonically organized as tetramers. MCD catalyzes the unprecedented oxidation of an α-methyl branched dicarboxylic acid CoA thioester. Substrate specificity is achieved by a cluster of three arginines that accommodates the terminal carboxyl group and a dedicated cavity that facilitates binding of the C2 methyl branch. MCD apparently evolved toward preventing the nonspecific oxidation of succinyl-CoA, which is a close structural homolog of (2S)-methylsuccinyl-CoA and an essential intermediate in central carbon metabolism. For different metabolic engineering and biotechnological applications, however, an enzyme that can oxidize succinyl-CoA to fumaryl-CoA is sought after. Based on the MCD structure, we were able to shift substrate specificity of MCD toward succinyl-CoA through active-site mutagenesis.
Keywords: X-ray crystallography; acyl-CoA dehydrogenase; bioengineering; enzyme catalysis; enzyme kinetics; enzyme mutation; enzyme purification; enzyme structure; ethylmalonyl-CoA pathway; flavin adenine dinucleotide (FAD); glutaryl-CoA dehydrogenase; isobutyryl-CoA dehydrogenase; mesaconyl-CoA.
© 2018 by The American Society for Biochemistry and Molecular Biology, Inc.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures
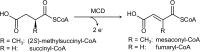
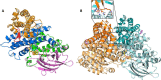


Similar articles
-
(2S)-Methylsuccinyl-CoA dehydrogenase closes the ethylmalonyl-CoA pathway for acetyl-CoA assimilation.Mol Microbiol. 2009 Sep;73(6):992-1008. doi: 10.1111/j.1365-2958.2009.06837.x. Epub 2009 Aug 23. Mol Microbiol. 2009. PMID: 19703103
-
Molecular Basis for Converting (2S)-Methylsuccinyl-CoA Dehydrogenase into an Oxidase.Molecules. 2017 Dec 28;23(1):68. doi: 10.3390/molecules23010068. Molecules. 2017. PMID: 29283425 Free PMC article.
-
Three-dimensional structure of the flavoenzyme acyl-CoA oxidase-II from rat liver, the peroxisomal counterpart of mitochondrial acyl-CoA dehydrogenase.J Biochem. 2002 Mar;131(3):365-74. doi: 10.1093/oxfordjournals.jbchem.a003111. J Biochem. 2002. PMID: 11872165
-
Biotechnological potential of the ethylmalonyl-CoA pathway.Appl Microbiol Biotechnol. 2011 Jan;89(1):17-25. doi: 10.1007/s00253-010-2873-z. Epub 2010 Sep 30. Appl Microbiol Biotechnol. 2011. PMID: 20882276 Review.
-
Novel B(12)-dependent acyl-CoA mutases and their biotechnological potential.Biochemistry. 2012 Aug 7;51(31):6039-46. doi: 10.1021/bi300827v. Epub 2012 Jul 23. Biochemistry. 2012. PMID: 22803641 Free PMC article. Review.
Cited by
-
Back to the future: Why we need enzymology to build a synthetic metabolism of the future.Beilstein J Org Chem. 2019 Feb 26;15:551-557. doi: 10.3762/bjoc.15.49. eCollection 2019. Beilstein J Org Chem. 2019. PMID: 30873239 Free PMC article. Review.
-
Structural basis for the broad substrate specificity of two acyl-CoA dehydrogenases FadE5 from mycobacteria.Proc Natl Acad Sci U S A. 2020 Jul 14;117(28):16324-16332. doi: 10.1073/pnas.2002835117. Epub 2020 Jun 29. Proc Natl Acad Sci U S A. 2020. PMID: 32601219 Free PMC article.
-
Pseudomonadal itaconate degradation gene cluster encodes enzymes for methylsuccinate utilization.Commun Biol. 2025 Jul 24;8(1):1099. doi: 10.1038/s42003-025-08538-2. Commun Biol. 2025. PMID: 40707647 Free PMC article.
References
-
- Ikeda Y., Dabrowski C., and Tanaka K. (1983) Separation and properties of five distinct acyl-CoA dehydrogenases from rat liver mitochondria: identification of a new 2-methyl branched chain acyl-CoA dehydrogenase. J. Biol. Chem. 258, 1066–1076 - PubMed
-
- Izai K., Uchida Y., Orii T., Yamamoto S., and Hashimoto T. (1992) Novel fatty acid β-oxidation enzymes in rat liver mitochondria. I. Purification and properties of very-long-chain acyl-coenzyme A dehydrogenase. J. Biol. Chem. 267, 1027–1033 - PubMed
-
- Nguyen T. V., Andresen B. S., Corydon T. J., Ghisla S., Abd-El Razik N., Mohsen A. W., Cederbaum S. D., Roe D. S., Roe C. R., Lench N. J., and Vockley J. (2002) Identification of isobutyryl-CoA dehydrogenase and its deficiency in humans. Mol. Genet. Metab. 77, 68–79 10.1016/S1096-7192(02)00152-X - DOI - PubMed
-
- Besrat A., Polan C. E., and Henderson L. M. (1969) Mammalian metabolism of glutaric acid. J. Biol. Chem. 244, 1461–1467 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

