Heat stress promotes longevity in budding yeast by relaxing the confinement of age-promoting factors in the mother cell
- PMID: 29283340
- PMCID: PMC5771669
- DOI: 10.7554/eLife.28329
Heat stress promotes longevity in budding yeast by relaxing the confinement of age-promoting factors in the mother cell
Abstract
Although individuals of many species inexorably age, a number of observations established that the rate of aging is modulated in response to a variety of mild stresses. Here, we investigated how heat stress promotes longevity in yeast. We show that upon growth at higher temperature, yeast cells relax the retention of DNA circles, which act as aging factors in the mother cell. The enhanced frequency at which circles redistribute to daughter cells was not due to changes of anaphase duration or nuclear shape but solely to the downregulation of the diffusion barrier in the nuclear envelope. This effect depended on the PKA and Tor1 pathways, downstream of stress-response kinase Pkc1. Inhibition of these responses restored barrier function and circle retention and abrogated the effect of heat stress on longevity. Our data indicate that redistribution of aging factors from aged cells to their progeny can be a mechanism for modulating longevity.
Keywords: S. cerevisiae; age confinement; aging; cell biology; heat stress; longevity.
Conflict of interest statement
No competing interests declared.
Figures







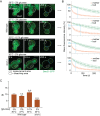





References
-
- Bolognesi A, Sliwa-Gonzalez A, Prasad R, Barral Y. fluorescence recovery after photo-bleaching (frap) and fluorescence loss in photo-bleaching (FLIP) experiments to study protein dynamics during budding yeast cell division. Methods in Molecular Biology. 2016;1369:25–44. doi: 10.1007/978-1-4939-3145-3_3. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

