LSD1 activation promotes inducible EMT programs and modulates the tumour microenvironment in breast cancer
- PMID: 29311580
- PMCID: PMC5758711
- DOI: 10.1038/s41598-017-17913-x
LSD1 activation promotes inducible EMT programs and modulates the tumour microenvironment in breast cancer
Erratum in
-
Author Correction: LSD1 activation promotes inducible EMT programs and modulates the tumour microenvironment in breast cancer.Sci Rep. 2019 Dec 5;9(1):18771. doi: 10.1038/s41598-019-55020-1. Sci Rep. 2019. PMID: 31801957 Free PMC article.
Abstract
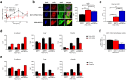
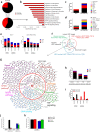
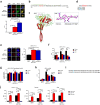
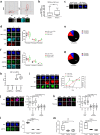
Complex regulatory networks control epithelial-to-mesenchymal transition (EMT) but the underlying epigenetic control is poorly understood. Lysine-specific demethylase 1 (LSD1) is a key histone demethylase that alters the epigenetic landscape. Here we explored the role of LSD1 in global epigenetic regulation of EMT, cancer stem cells (CSCs), the tumour microenvironment, and therapeutic resistance in breast cancer. LSD1 induced pan-genomic gene expression in networks implicated in EMT and selectively elicits gene expression programs in CSCs whilst repressing non-CSC programs. LSD1 phosphorylation at serine-111 (LSD1-s111p) by chromatin anchored protein kinase C-theta (PKC-θ), is critical for its demethylase and EMT promoting activity and LSD1-s111p is enriched in chemoresistant cells in vivo. LSD1 couples to PKC-θ on the mesenchymal gene epigenetic template promotes LSD1-mediated gene induction. In vivo, chemotherapy reduced tumour volume, and when combined with an LSD1 inhibitor, abrogated the mesenchymal signature and promoted an innate, M1 macrophage-like tumouricidal immune response. Circulating tumour cells (CTCs) from metastatic breast cancer (MBC) patients were enriched with LSD1 and pharmacological blockade of LSD1 suppressed the mesenchymal and stem-like signature in these patient-derived CTCs. Overall, LSD1 inhibition may serve as a promising epigenetic adjuvant therapy to subvert its pleiotropic roles in breast cancer progression and treatment resistance.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures



Similar articles
-
The malignant brain tumor (MBT) domain protein SFMBT1 is an integral histone reader subunit of the LSD1 demethylase complex for chromatin association and epithelial-to-mesenchymal transition.J Biol Chem. 2013 Sep 20;288(38):27680-27691. doi: 10.1074/jbc.M113.482349. Epub 2013 Aug 8. J Biol Chem. 2013. PMID: 23928305 Free PMC article.
-
HOTAIR lncRNA promotes epithelial-mesenchymal transition by redistributing LSD1 at regulatory chromatin regions.EMBO Rep. 2021 Jul 5;22(7):e50193. doi: 10.15252/embr.202050193. Epub 2021 May 6. EMBO Rep. 2021. PMID: 33960111 Free PMC article.
-
Lysine-Specific Demethylase 1 Mediates AKT Activity and Promotes Epithelial-to-Mesenchymal Transition in PIK3CA-Mutant Colorectal Cancer.Mol Cancer Res. 2020 Feb;18(2):264-277. doi: 10.1158/1541-7786.MCR-19-0748. Epub 2019 Nov 8. Mol Cancer Res. 2020. PMID: 31704733 Free PMC article.
-
Epigenetic regulation of epithelial to mesenchymal transition by the Lysine-specific demethylase LSD1/KDM1A.Biochim Biophys Acta Gene Regul Mech. 2017 Sep;1860(9):905-910. doi: 10.1016/j.bbagrm.2017.07.001. Epub 2017 Jul 15. Biochim Biophys Acta Gene Regul Mech. 2017. PMID: 28720390 Review.
-
Epigenetic Regulation of Inflammatory Cytokine-Induced Epithelial-To-Mesenchymal Cell Transition and Cancer Stem Cell Generation.Cells. 2019 Sep 25;8(10):1143. doi: 10.3390/cells8101143. Cells. 2019. PMID: 31557902 Free PMC article. Review.
Cited by
-
Targeting epigenetic regulators to overcome drug resistance in cancers.Signal Transduct Target Ther. 2023 Feb 17;8(1):69. doi: 10.1038/s41392-023-01341-7. Signal Transduct Target Ther. 2023. PMID: 36797239 Free PMC article. Review.
-
Lysine-Specific Histone Demethylase 1A Regulates Macrophage Polarization and Checkpoint Molecules in the Tumor Microenvironment of Triple-Negative Breast Cancer.Front Immunol. 2019 Jun 12;10:1351. doi: 10.3389/fimmu.2019.01351. eCollection 2019. Front Immunol. 2019. PMID: 31249575 Free PMC article.
-
Advances in liquid biopsy using circulating tumor cells and circulating cell-free tumor DNA for detection and monitoring of breast cancer.Clin Exp Med. 2019 Aug;19(3):271-279. doi: 10.1007/s10238-019-00563-w. Epub 2019 Jun 12. Clin Exp Med. 2019. PMID: 31190187 Review.
-
Nuclear-Biased DUSP6 Expression is Associated with Cancer Spreading Including Brain Metastasis in Triple-Negative Breast Cancer.Int J Mol Sci. 2019 Jun 24;20(12):3080. doi: 10.3390/ijms20123080. Int J Mol Sci. 2019. PMID: 31238530 Free PMC article.
-
Histone modifications in drug-resistant cancers: From a cancer stem cell and immune evasion perspective.Exp Mol Med. 2023 Jul;55(7):1333-1347. doi: 10.1038/s12276-023-01014-z. Epub 2023 Jul 3. Exp Mol Med. 2023. PMID: 37394580 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

