Identifying Rodent Resting-State Brain Networks with Independent Component Analysis
- PMID: 29311770
- PMCID: PMC5733053
- DOI: 10.3389/fnins.2017.00685
Identifying Rodent Resting-State Brain Networks with Independent Component Analysis
Abstract
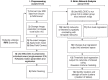
Rodent models have opened the door to a better understanding of the neurobiology of brain disorders and increased our ability to evaluate novel treatments. Resting-state functional magnetic resonance imaging (rs-fMRI) allows for in vivo exploration of large-scale brain networks with high spatial resolution. Its application in rodents affords researchers a powerful translational tool to directly assess/explore the effects of various pharmacological, lesion, and/or disease states on known neural circuits within highly controlled settings. Integration of animal and human research at the molecular-, systems-, and behavioral-levels using diverse neuroimaging techniques empowers more robust interrogations of abnormal/ pathological processes, critical for evolving our understanding of neuroscience. We present a comprehensive protocol to evaluate resting-state brain networks using Independent Component Analysis (ICA) in rodent model. Specifically, we begin with a brief review of the physiological basis for rs-fMRI technique and overview of rs-fMRI studies in rodents to date, following which we provide a robust step-by-step approach for rs-fMRI investigation including data collection, computational preprocessing, and brain network analysis. Pipelines are interwoven with underlying theory behind each step and summarized methodological considerations, such as alternative methods available and current consensus in the literature for optimal results. The presented protocol is designed in such a way that investigators without previous knowledge in the field can implement the analysis and obtain viable results that reliably detect significant differences in functional connectivity between experimental groups. Our goal is to empower researchers to implement rs-fMRI in their respective fields by incorporating technical considerations to date into a workable methodological framework.
Keywords: BOLD; ICA; MRI; fMRI; protocol; resting-state networks; review; rs-fMRI.
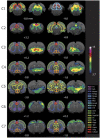
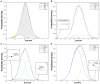
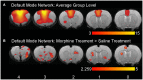
Figures



References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

