Hedgehog Pathway Drives Fusion-Negative Rhabdomyosarcoma Initiated From Non-myogenic Endothelial Progenitors
- PMID: 29316425
- PMCID: PMC5790179
- DOI: 10.1016/j.ccell.2017.12.001
Hedgehog Pathway Drives Fusion-Negative Rhabdomyosarcoma Initiated From Non-myogenic Endothelial Progenitors
Abstract
Rhabdomyosarcoma (RMS) is a pediatric soft tissue sarcoma that histologically resembles embryonic skeletal muscle. RMS occurs throughout the body and an exclusively myogenic origin does not account for RMS occurring in sites devoid of skeletal muscle. We previously described an RMS model activating a conditional constitutively active Smoothened mutant (SmoM2) with aP2-Cre. Using genetic fate mapping, we show SmoM2 expression in Cre-expressing endothelial progenitors results in myogenic transdifferentiation and RMS. We show that endothelium and skeletal muscle within the head and neck arise from Kdr-expressing progenitors, and that hedgehog pathway activation results in aberrant expression of myogenic specification factors as a potential mechanism driving RMS genesis. These findings suggest that RMS can originate from aberrant development of non-myogenic cells.
Keywords: Tbx1; endothelium; hedgehog; myogenesis; rhabdomyosarcoma; sarcoma; skeletal muscle; transdifferentiation.
Copyright © 2017 Elsevier Inc. All rights reserved.
Figures


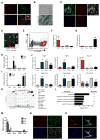




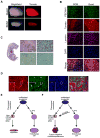
Comment in
-
What's in a Name? Cell Fate Reprogramming in Sarcomagenesis.Cancer Cell. 2018 Jan 8;33(1):5-7. doi: 10.1016/j.ccell.2017.12.005. Cancer Cell. 2018. PMID: 29316432
-
Fusion-Negative Rhabdomyosarcoma Can Arise from Endothelial Cells.Cancer Discov. 2018 Mar;8(3):261. doi: 10.1158/2159-8290.CD-RW2018-009. Epub 2018 Jan 19. Cancer Discov. 2018. PMID: 29352048
References
-
- Benjamini Y, Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society Series B (Methodological) 1995;57:289–300.
-
- Borycki AG, Brunk B, Tajbakhsh S, Buckingham M, Chiang C, Emerson CP., Jr Sonic hedgehog controls epaxial muscle determination through Myf5 activation. Development. 1999;126:4053–4063. - PubMed
-
- Bridge JA, Liu J, Weibolt V, Baker KS, Perry D, Kruger R, Qualman S, Barr F, Sorensen P, Triche T, Suijkerbuijk R. Novel genomic imbalances in embryonal rhabdomyosarcoma revealed by comparative genomic hybridization and fluorescence in situ hybridization: an intergroup rhabdomyosarcoma study. Genes, chromosomes & cancer. 2000;27:337–344. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

