Germline and somatic variant identification using BGISEQ-500 and HiSeq X Ten whole genome sequencing
- PMID: 29320538
- PMCID: PMC5761881
- DOI: 10.1371/journal.pone.0190264
Germline and somatic variant identification using BGISEQ-500 and HiSeq X Ten whole genome sequencing
Abstract
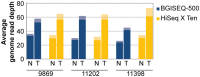
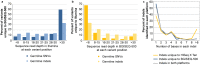
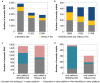
Technological innovation and increased affordability have contributed to the widespread adoption of genome sequencing technologies in biomedical research. In particular large cancer research consortia have embraced next generation sequencing, and have used the technology to define the somatic mutation landscape of multiple cancer types. These studies have primarily utilised the Illumina HiSeq platforms. In this study we performed whole genome sequencing of three malignant pleural mesothelioma and matched normal samples using a new platform, the BGISEQ-500, and compared the results obtained with Illumina HiSeq X Ten. Germline and somatic, single nucleotide variants and small insertions or deletions were independently identified from data aligned human genome reference. The BGISEQ-500 and HiSeq X Ten platforms showed high concordance for germline calls with genotypes from SNP arrays (>99%). The germline and somatic single nucleotide variants identified in both sequencing platforms were highly concordant (86% and 72% respectively). These results indicate the potential applicability of the BGISEQ-500 platform for the identification of somatic and germline single nucleotide variants by whole genome sequencing. The BGISEQ-500 datasets described here represent the first publicly-available cancer genome sequencing performed using this platform.
Conflict of interest statement
Figures

Similar articles
-
Comparison and evaluation of two exome capture kits and sequencing platforms for variant calling.BMC Genomics. 2015 Aug 5;16(1):581. doi: 10.1186/s12864-015-1796-6. BMC Genomics. 2015. PMID: 26242175 Free PMC article.
-
Comparative analysis of novel MGISEQ-2000 sequencing platform vs Illumina HiSeq 2500 for whole-genome sequencing.PLoS One. 2020 Mar 16;15(3):e0230301. doi: 10.1371/journal.pone.0230301. eCollection 2020. PLoS One. 2020. PMID: 32176719 Free PMC article.
-
A new massively parallel nanoball sequencing platform for whole exome research.BMC Bioinformatics. 2019 Mar 25;20(1):153. doi: 10.1186/s12859-019-2751-3. BMC Bioinformatics. 2019. PMID: 30909888 Free PMC article.
-
Whole Genome Library Construction for Next Generation Sequencing.Methods Mol Biol. 2018;1706:151-161. doi: 10.1007/978-1-4939-7471-9_8. Methods Mol Biol. 2018. PMID: 29423797 Review.
-
Direct mutation analysis by high-throughput sequencing: from germline to low-abundant, somatic variants.Mutat Res. 2012 Jan 3;729(1-2):1-15. doi: 10.1016/j.mrfmmm.2011.10.001. Epub 2011 Oct 12. Mutat Res. 2012. PMID: 22016070 Free PMC article. Review.
Cited by
-
Genome sequencing of human in vitro fertilisation embryos for pathogenic variation screening.Sci Rep. 2020 Mar 2;10(1):3795. doi: 10.1038/s41598-020-60704-0. Sci Rep. 2020. PMID: 32123222 Free PMC article.
-
Comparison of the DNBSEQ platform and Illumina HiSeq 2000 for bacterial genome assembly.Sci Rep. 2024 Jan 14;14(1):1292. doi: 10.1038/s41598-024-51725-0. Sci Rep. 2024. PMID: 38221534 Free PMC article.
-
The application of NIPT using combinatorial probe-anchor synthesis to identify sex chromosomal aneuploidies (SCAs) in a cohort of 570 pregnancies.Mol Cytogenet. 2018 Dec 3;11:59. doi: 10.1186/s13039-018-0407-z. eCollection 2018. Mol Cytogenet. 2018. PMID: 30524505 Free PMC article.
-
Assessing reproducibility of inherited variants detected with short-read whole genome sequencing.Genome Biol. 2022 Jan 3;23(1):2. doi: 10.1186/s13059-021-02569-8. Genome Biol. 2022. PMID: 34980216 Free PMC article.
-
SEQdata-BEACON: a comprehensive database of sequencing performance and statistical tools for performance evaluation and yield simulation in BGISEQ-500.BioData Min. 2019 Nov 15;12:21. doi: 10.1186/s13040-019-0209-9. eCollection 2019. BioData Min. 2019. PMID: 31807141 Free PMC article.
References
-
- Goodwin S, McPherson JD, McCombie WR. Coming of age: ten years of next-generation sequencing technologies. Nat Rev Genet. 2016;17(6):333–51. doi: 10.1038/nrg.2016.49 . - DOI - PMC - PubMed
-
- Mardis ER. A decade’s perspective on DNA sequencing technology. Nature. 2011;470(7333):198–203. doi: 10.1038/nature09796 . - DOI - PubMed
-
- Cancer Genome Atlas Research N, Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA, et al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Genet. 2013;45(10):1113–20. doi: 10.1038/ng.2764 . - DOI - PMC - PubMed
-
- International Cancer Genome C, Hudson TJ, Anderson W, Artez A, Barker AD, Bell C, et al. International network of cancer genome projects. Nature. 2010;464(7291):993–8. doi: 10.1038/nature08987 . - DOI - PMC - PubMed
-
- Alexandrov LB, Nik-Zainal S, Wedge DC, Campbell PJ, Stratton MR. Deciphering signatures of mutational processes operative in human cancer. Cell reports. 2013;3(1):246–59. doi: 10.1016/j.celrep.2012.12.008 . - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

