Structural Basis for Activity and Specificity of an Anticoagulant Anti-FXIa Monoclonal Antibody and a Reversal Agent
- PMID: 29336885
- PMCID: PMC5803430
- DOI: 10.1016/j.str.2017.12.010
Structural Basis for Activity and Specificity of an Anticoagulant Anti-FXIa Monoclonal Antibody and a Reversal Agent
Abstract
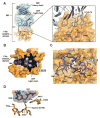
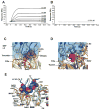
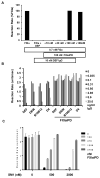
Coagulation factor XIa is a candidate target for anticoagulants that better separate antithrombotic efficacy from bleeding risk. We report a co-crystal structure of the FXIa protease domain with DEF, a human monoclonal antibody that blocks FXIa function and prevents thrombosis in animal models without detectable increased bleeding. The light chain of DEF occludes the FXIa S1 subsite and active site, while the heavy chain provides electrostatic interactions with the surface of FXIa. The structure accounts for the specificity of DEF for FXIa over its zymogen and related proteases, its active-site-dependent binding, and its ability to inhibit substrate cleavage. The inactive FXIa protease domain used to obtain the DEF-FXIa crystal structure reversed anticoagulant activity of DEF in plasma and in vivo and the activity of a small-molecule FXIa active-site inhibitor in vitro. DEF and this reversal agent for FXIa active-site inhibitors may help support clinical development of FXIa-targeting anticoagulants.
Keywords: IgG; active-site inhibitor; anticoagulant; coagulation; crystal structure; factor XI; hemostasis; intrinsic pathway; protease inhibitor; protease-blocking antibody; thrombosis.
Copyright © 2017 Elsevier Ltd. All rights reserved.
Figures


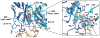
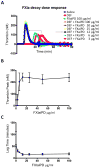

References
-
- Asakai R, Chung DW, Davie EW, Seligsohn U. Factor XI deficiency in Ashkenazi Jews in Israel. The New England journal of medicine. 1991;325:153–158. - PubMed
-
- Bi L, Lawler AM, Antonarakis SE, High KA, Gearhart JD, Kazazian HH., Jr Targeted disruption of the mouse factor VIII gene produces a model of haemophilia A [letter] Nature genetics. 1995;10:119–121. - PubMed
-
- Bugge TH, Xiao Q, Kombrinck KW, Flick MJ, Holmbäck K, Danton MJ, Colbert MC, Witte DP, Fujikawa K, Davie EW, et al. Fatal embryonic bleeding events in mice lacking tissue factor, the cell-associated initiator of blood coagulation. Proceedings of the National Academy of Sciences of the United States of America. 1996;93:6258–6263. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

