A Non-canonical BCOR-PRC1.1 Complex Represses Differentiation Programs in Human ESCs
- PMID: 29337181
- PMCID: PMC5797497
- DOI: 10.1016/j.stem.2017.12.002
A Non-canonical BCOR-PRC1.1 Complex Represses Differentiation Programs in Human ESCs
Abstract
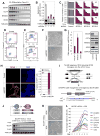
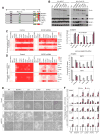
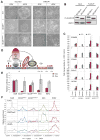
Polycomb group proteins regulate self-renewal and differentiation in many stem cell systems. When assembled into two canonical complexes, PRC1 and PRC2, they sequentially deposit H3K27me3 and H2AK119ub histone marks and establish repressive chromatin, referred to as Polycomb domains. Non-canonical PRC1 complexes retain RING1/RNF2 E3-ubiquitin ligases but have unique sets of accessory subunits. How these non-canonical complexes recognize and regulate their gene targets remains poorly understood. Here, we show that the BCL6 co-repressor (BCOR), a member of the PRC1.1 complex, is critical for maintaining primed pluripotency in human embryonic stem cells (ESCs). BCOR depletion leads to the erosion of Polycomb domains at key developmental loci and the initiation of differentiation along endoderm and mesoderm lineages. The C terminus of BCOR regulates the assembly and targeting of the PRC1.1 complex, while the N terminus contributes to BCOR-PRC1.1 repressor function. Our findings advance understanding of Polycomb targeting and repression in ESCs and could apply broadly across developmental systems.
Keywords: BCOR; human embryonic stem cells; pluripotency; polycomb repressive complexes.
Copyright © 2017 Elsevier Inc. All rights reserved.
Figures




References
-
- Adamo A, Sese B, Boue S, Castano J, Paramonov I, Barrero MJ, Izpisua Belmonte JC. LSD1 regulates the balance between self-renewal and differentiation in human embryonic stem cells. Nature cell biology. 2011;13:652–659. - PubMed
-
- Beguelin W, Teater M, Gearhart MD, Calvo Fernandez MT, Goldstein RL, Cardenas MG, Hatzi K, Rosen M, Shen H, Corcoran CM, et al. EZH2 and BCL6 Cooperate to Assemble CBX8-BCOR Complex to Repress Bivalent Promoters, Mediate Germinal Center Formation and Lymphomagenesis. Cancer cell. 2016;30:197–213. - PMC - PubMed
-
- Boulard M, Edwards JR, Bestor TH. FBXL10 protects Polycomb-bound genes from hypermethylation. Nature genetics. 2015;47:479–485. - PubMed
-
- Boyer LA, Plath K, Zeitlinger J, Brambrink T, Medeiros LA, Lee TI, Levine SS, Wernig M, Tajonar A, Ray MK, et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature. 2006;441:349–353. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

