Hot-spot KIF5A mutations cause familial ALS
- PMID: 29342275
- PMCID: PMC5837483
- DOI: 10.1093/brain/awx370
Hot-spot KIF5A mutations cause familial ALS
Abstract
Heterozygous missense mutations in the N-terminal motor or coiled-coil domains of the kinesin family member 5A (KIF5A) gene cause monogenic spastic paraplegia (HSP10) and Charcot-Marie-Tooth disease type 2 (CMT2). Moreover, heterozygous de novo frame-shift mutations in the C-terminal domain of KIF5A are associated with neonatal intractable myoclonus, a neurodevelopmental syndrome. These findings, together with the observation that many of the disease genes associated with amyotrophic lateral sclerosis disrupt cytoskeletal function and intracellular transport, led us to hypothesize that mutations in KIF5A are also a cause of amyotrophic lateral sclerosis. Using whole exome sequencing followed by rare variant analysis of 426 patients with familial amyotrophic lateral sclerosis and 6137 control subjects, we detected an enrichment of KIF5A splice-site mutations in amyotrophic lateral sclerosis (2/426 compared to 0/6137 in controls; P = 4.2 × 10-3), both located in a hot-spot in the C-terminus of the protein and predicted to affect splicing exon 27. We additionally show co-segregation with amyotrophic lateral sclerosis of two canonical splice-site mutations in two families. Investigation of lymphoblast cell lines from patients with KIF5A splice-site mutations revealed the loss of mutant RNA expression and suggested haploinsufficiency as the most probable underlying molecular mechanism. Furthermore, mRNA sequencing of a rare non-synonymous missense mutation (predicting p.Arg1007Gly) located in the C-terminus of the protein shortly upstream of the splice donor of exon 27 revealed defective KIF5A pre-mRNA splicing in respective patient-derived cell lines owing to abrogation of the donor site. Finally, the non-synonymous single nucleotide variant rs113247976 (minor allele frequency = 1.00% in controls, n = 6137), also located in the C-terminal region [p.(Pro986Leu) in exon 26], was significantly enriched in familial amyotrophic lateral sclerosis patients (minor allele frequency = 3.40%; P = 1.28 × 10-7). Our study demonstrates that mutations located specifically in a C-terminal hotspot of KIF5A can cause a classical amyotrophic lateral sclerosis phenotype, and underline the involvement of intracellular transport processes in amyotrophic lateral sclerosis pathogenesis.
Figures
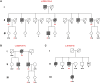


Comment in
-
Adult-onset distal spinal muscular atrophy: a new phenotype associated with KIF5A mutations.Brain. 2019 Dec 1;142(12):e66. doi: 10.1093/brain/awz317. Brain. 2019. PMID: 31612903 No abstract available.
References
-
- Andersen PM, Borasio GD, Dengler R, Hardiman O, Kollewe K, Leigh PN, et al. EFNS task force on management of amyotrophic lateral sclerosis: guidelines for diagnosing and clinical care of patients and relatives. An evidence-based review with good practice points. Eur J Neurol 2005; 12: 921–38. - PubMed
-
- Andersen PM, Abrahams S, Borasio GD, de Carvalho M, Chio A, Van Damme P, et al. EFNS guidelines on the Clinical Management of Amyotrophic Lateral Sclerosis (MALS)—revised report of an EFNS task force. Eur J Neurol 2012; 19: 360–75. - PubMed
-
- Brenner D, Müller K, Wieland T, Weydt P, Böhm S, Lulé D, et al. NEK1 mutations in familial amyotrophic lateral sclerosis. Brain 2016; 139: e28. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

