Ceratocystis cacaofunesta genome analysis reveals a large expansion of extracellular phosphatidylinositol-specific phospholipase-C genes (PI-PLC)
- PMID: 29343217
- PMCID: PMC5773145
- DOI: 10.1186/s12864-018-4440-4
Ceratocystis cacaofunesta genome analysis reveals a large expansion of extracellular phosphatidylinositol-specific phospholipase-C genes (PI-PLC)
Abstract
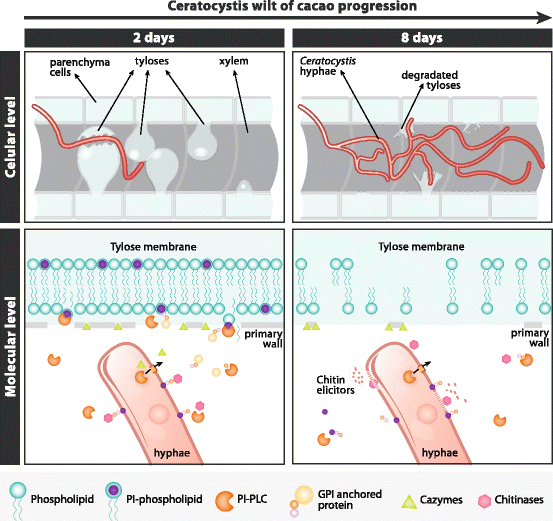
Background: The Ceratocystis genus harbors a large number of phytopathogenic fungi that cause xylem parenchyma degradation and vascular destruction on a broad range of economically important plants. Ceratocystis cacaofunesta is a necrotrophic fungus responsible for lethal wilt disease in cacao. The aim of this work is to analyze the genome of C. cacaofunesta through a comparative approach with genomes of other Sordariomycetes in order to better understand the molecular basis of pathogenicity in the Ceratocystis genus.
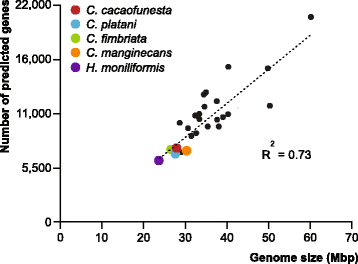
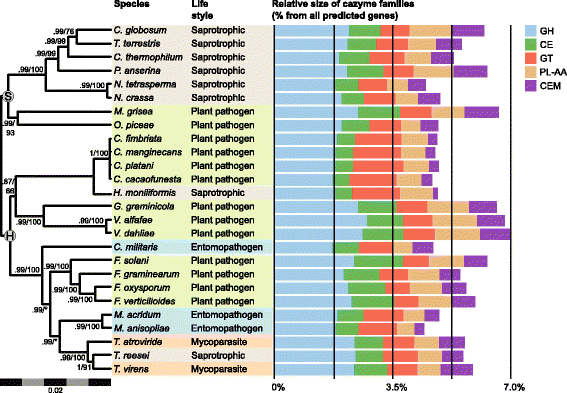
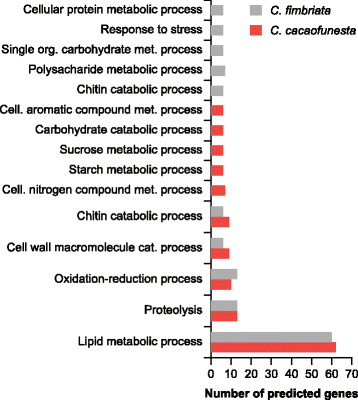
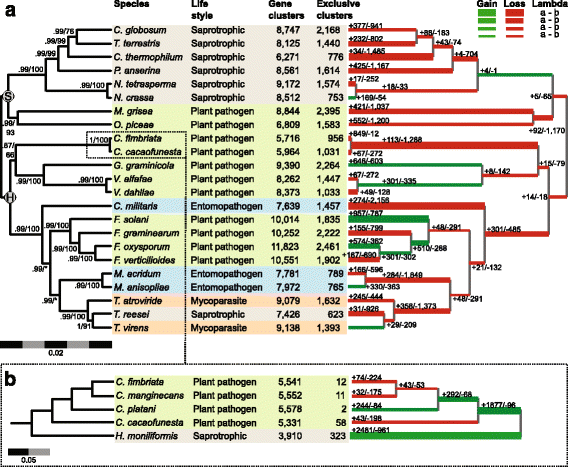
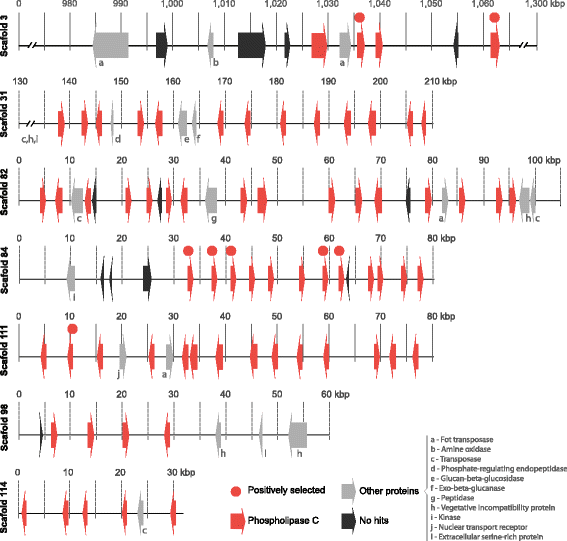
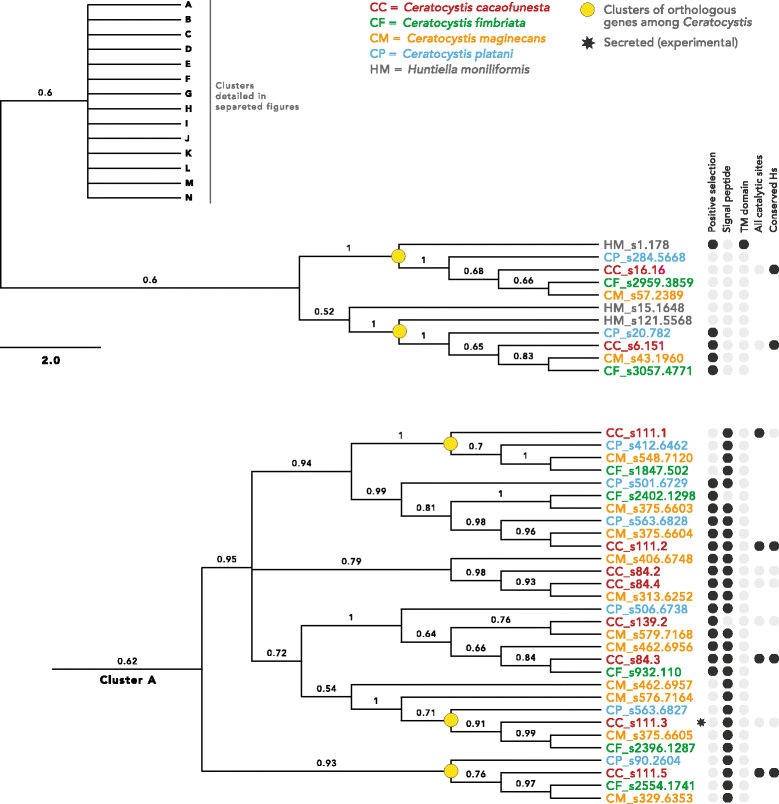
Results: We present an analysis of the C. cacaofunesta genome focusing on secreted proteins that might constitute pathogenicity factors. Comparative genome analyses among five Ceratocystidaceae species and 23 other Sordariomycetes fungi showed a strong reduction in gene content of the Ceratocystis genus. However, some gene families displayed a remarkable expansion, in particular, the Phosphatidylinositol specific phospholipases-C (PI-PLC) family. Also, evolutionary rate calculations suggest that the evolution process of this family was guided by positive selection. Interestingly, among the 82 PI-PLCs genes identified in the C. cacaofunesta genome, 70 genes encoding extracellular PI-PLCs are grouped in eight small scaffolds surrounded by transposon fragments and scars that could be involved in the rapid evolution of the PI-PLC family. Experimental secretome using LC-MS/MS validated 24% (86 proteins) of the total predicted secretome (342 proteins), including four PI-PLCs and other important pathogenicity factors.
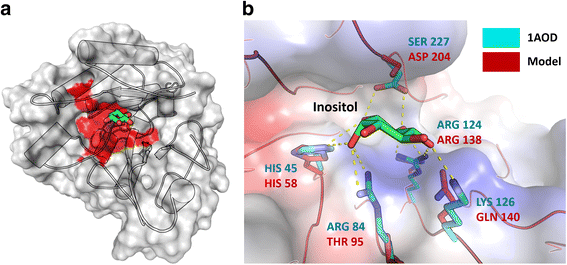
Conclusion: Analysis of the Ceratocystis cacaofunesta genome provides evidence that PI-PLCs may play a role in pathogenicity. Subsequent functional studies will be aimed at evaluating this hypothesis. The observed genetic arsenals, together with the analysis of the PI-PLC family shown in this work, reveal significant differences in the Ceratocystis genome compared to the classical vascular fungi, Verticillium and Fusarium. Altogether, our analyses provide new insights into the evolution and the molecular basis of plant pathogenicity.
Keywords: Ceratocystis wilt of cacao; Phosphoinositide-specific phospholipases C (PI PLC); Plant pathogen.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures

Similar articles
-
A repertoire of candidate effector proteins of the fungus Ceratocystis cacaofunesta.Sci Rep. 2023 Sep 29;13(1):16368. doi: 10.1038/s41598-023-43117-7. Sci Rep. 2023. PMID: 37773261 Free PMC article.
-
Global analyses of Ceratocystis cacaofunesta mitochondria: from genome to proteome.BMC Genomics. 2013 Feb 11;14:91. doi: 10.1186/1471-2164-14-91. BMC Genomics. 2013. PMID: 23394930 Free PMC article.
-
Volatile Organic Compounds from Ceratocystis cacaofunesta, a Causal Agent of Ceratocystis Wilt of Cacao.J Chem Ecol. 2024 Nov;50(11):807-814. doi: 10.1007/s10886-024-01542-6. Epub 2024 Aug 27. J Chem Ecol. 2024. PMID: 39190193
-
Plant phosphoinositide-dependent phospholipases C: variations around a canonical theme.Biochimie. 2014 Jan;96:144-57. doi: 10.1016/j.biochi.2013.07.004. Epub 2013 Jul 12. Biochimie. 2014. PMID: 23856562 Review.
-
Phosphoinositide-specific phospholipase C in health and disease.J Lipid Res. 2015 Oct;56(10):1853-60. doi: 10.1194/jlr.R057984. Epub 2015 Mar 27. J Lipid Res. 2015. PMID: 25821234 Free PMC article. Review.
Cited by
-
Uncovering a Complex Virome Associated with the Cacao Pathogens Ceratocystis cacaofunesta and Ceratocystis fimbriata.Pathogens. 2023 Feb 9;12(2):287. doi: 10.3390/pathogens12020287. Pathogens. 2023. PMID: 36839559 Free PMC article.
-
Whole genome sequencing and analysis of multiple isolates of Ceratocystis destructans, the causal agent of Ceratocystis canker of almond in California.Sci Rep. 2023 Sep 8;13(1):14873. doi: 10.1038/s41598-023-41746-6. Sci Rep. 2023. PMID: 37684350 Free PMC article.
-
Phospholipase C: Diverse functions in plant biotic stress resistance and fungal pathogenicity.Mol Plant Pathol. 2023 Sep;24(9):1192-1202. doi: 10.1111/mpp.13343. Epub 2023 Apr 29. Mol Plant Pathol. 2023. PMID: 37119461 Free PMC article. Review.
-
Genome-Wide Identification and Expression Profile Analysis of the Phospholipase C Gene Family in Wheat (Triticum aestivum L.).Plants (Basel). 2020 Jul 13;9(7):885. doi: 10.3390/plants9070885. Plants (Basel). 2020. PMID: 32668812 Free PMC article.
-
A repertoire of candidate effector proteins of the fungus Ceratocystis cacaofunesta.Sci Rep. 2023 Sep 29;13(1):16368. doi: 10.1038/s41598-023-43117-7. Sci Rep. 2023. PMID: 37773261 Free PMC article.
References
-
- Cabrera OG, et al. Ceratocystis wilt pathogens: history and biology—highlighting C. cacaofunesta, the causal agent of wilt disease of cacao. In: Bailey AB, Meinhardt WL, et al., editors. Cacao diseases: a history of old enemies and new encounters. Cham: Springer International Publishing; 2016. pp. 383–428.
Publication types
MeSH terms
Substances
Grants and funding
- 2009/50119-9/Fundação de Amparo à Pesquisa do Estado de São Paulo/International
- 2009/18467-7/Fundação de Amparo à Pesquisa do Estado de São Paulo/International
- 142357/2014-6/Conselho Nacional de Desenvolvimento Científico e Tecnológico/International
- 2013/08293-7/Center for Computational Engineering and Sciences - FAPESP/Cepid/International
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

