Compositional Bias in Naïve and Chemically-modified Phage-Displayed Libraries uncovered by Paired-end Deep Sequencing
- PMID: 29352178
- PMCID: PMC5775325
- DOI: 10.1038/s41598-018-19439-2
Compositional Bias in Naïve and Chemically-modified Phage-Displayed Libraries uncovered by Paired-end Deep Sequencing
Abstract
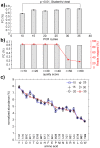
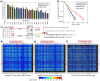
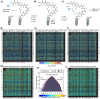
Understanding the composition of a genetically-encoded (GE) library is instrumental to the success of ligand discovery. In this manuscript, we investigate the bias in GE-libraries of linear, macrocyclic and chemically post-translationally modified (cPTM) tetrapeptides displayed on the M13KE platform, which are produced via trinucleotide cassette synthesis (19 codons) and NNK-randomized codon. Differential enrichment of synthetic DNA {S}, ligated vector {L} (extension and ligation of synthetic DNA into the vector), naïve libraries {N} (transformation of the ligated vector into the bacteria followed by expression of the library for 4.5 hours to yield a "naïve" library), and libraries chemically modified by aldehyde ligation and cysteine macrocyclization {M} characterized by paired-end deep sequencing, detected a significant drop in diversity in {L} → {N}, but only a minor compositional difference in {S} → {L} and {N} → {M}. Libraries expressed at the N-terminus of phage protein pIII censored positively charged amino acids Arg and Lys; libraries expressed between pIII domains N1 and N2 overcame Arg/Lys-censorship but introduced new bias towards Gly and Ser. Interrogation of biases arising from cPTM by aldehyde ligation and cysteine macrocyclization unveiled censorship of sequences with Ser/Phe. Analogous analysis can be used to explore library diversity in new display platforms and optimize cPTM of these libraries.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




Similar articles
-
Late-Stage Reshaping of Phage-Displayed Libraries to Macrocyclic and Bicyclic Landscapes using a Multipurpose Linchpin.J Am Chem Soc. 2025 Jan 8;147(1):789-800. doi: 10.1021/jacs.4c13561. Epub 2024 Dec 19. J Am Chem Soc. 2025. PMID: 39702930
-
Deep sequencing analysis of phage libraries using Illumina platform.Methods. 2012 Sep;58(1):47-55. doi: 10.1016/j.ymeth.2012.07.006. Epub 2012 Jul 20. Methods. 2012. PMID: 22819855
-
Silent Encoding of Chemical Post-Translational Modifications in Phage-Displayed Libraries.J Am Chem Soc. 2016 Jan 13;138(1):32-5. doi: 10.1021/jacs.5b10390. Epub 2015 Dec 28. J Am Chem Soc. 2016. PMID: 26683999
-
Next-generation sequencing of phage-displayed peptide libraries.Methods Mol Biol. 2015;1248:249-66. doi: 10.1007/978-1-4939-2020-4_17. Methods Mol Biol. 2015. PMID: 25616338 Review.
-
Biased selection of propagation-related TUPs from phage display peptide libraries.Amino Acids. 2017 Aug;49(8):1293-1308. doi: 10.1007/s00726-017-2452-z. Epub 2017 Jun 29. Amino Acids. 2017. PMID: 28664268 Review.
Cited by
-
Evolving a Peptide: Library Platforms and Diversification Strategies.Int J Mol Sci. 2019 Dec 27;21(1):215. doi: 10.3390/ijms21010215. Int J Mol Sci. 2019. PMID: 31892275 Free PMC article. Review.
-
Optimized Construction of a Yeast SICLOPPS Library for Unbiased In Vivo Selection of Cyclic Peptides.Biochemistry. 2024 Dec 17;63(24):3273-3286. doi: 10.1021/acs.biochem.4c00013. Epub 2024 Dec 6. Biochemistry. 2024. PMID: 39642937 Free PMC article.
-
Closing the loop with reactions at the N-terminus.Nat Rev Chem. 2025 Mar;9(3):142-143. doi: 10.1038/s41570-025-00697-4. Nat Rev Chem. 2025. PMID: 39953291 No abstract available.
-
Construction of Targeting-Peptide-Based Imaging Reagents and Their Application in Bioimaging.Chem Biomed Imaging. 2023 Dec 4;2(4):233-249. doi: 10.1021/cbmi.3c00104. eCollection 2024 Apr 22. Chem Biomed Imaging. 2023. PMID: 39473775 Free PMC article. Review.
-
Exploring the Post-translational Enzymology of PaaA by mRNA Display.J Am Chem Soc. 2020 Mar 18;142(11):5024-5028. doi: 10.1021/jacs.0c01576. Epub 2020 Mar 6. J Am Chem Soc. 2020. PMID: 32109054 Free PMC article.
References
-
- Lee YJ, et al. Fabricating genetically engineered high-power lithium-ion batteries using multiple virus genes. Science. 2009;324:1051–1055. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

