Compartmentalization of HP1 Proteins in Pluripotency Acquisition and Maintenance
- PMID: 29358085
- PMCID: PMC5830946
- DOI: 10.1016/j.stemcr.2017.12.016
Compartmentalization of HP1 Proteins in Pluripotency Acquisition and Maintenance
Abstract
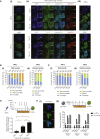
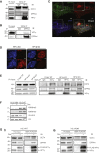
The heterochromatin protein 1 (HP1) family is involved in various functions with maintenance of chromatin structure. During murine somatic cell reprogramming, we find that early depletion of HP1γ reduces the generation of induced pluripotent stem cells, while late depletion enhances the process, with a concomitant change from a centromeric to nucleoplasmic localization and elongation-associated histone H3.3 enrichment. Depletion of heterochromatin anchoring protein SENP7 increased reprogramming efficiency to a similar extent as HP1γ, indicating the importance of HP1γ release from chromatin for pluripotency acquisition. HP1γ interacted with OCT4 and DPPA4 in HP1α and HP1β knockouts and in H3K9 methylation depleted H3K9M embryonic stem cell (ESC) lines. HP1α and HP1γ complexes in ESCs differed in association with histones, the histone chaperone CAF1 complex, and specific components of chromatin-modifying complexes such as DPY30, implying distinct functional contributions. Taken together, our results reveal the complex contribution of the HP1 proteins to pluripotency.
Keywords: Dppa4; H3.3; H3K9M; HP1α knockout; HP1β knockout; HP1γ knockout; Senp7; iPSC; pluripotency; reprogramming.
Copyright © 2017 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures





Similar articles
-
Proteomic and genomic approaches reveal critical functions of H3K9 methylation and heterochromatin protein-1γ in reprogramming to pluripotency.Nat Cell Biol. 2013 Jul;15(7):872-82. doi: 10.1038/ncb2768. Epub 2013 Jun 9. Nat Cell Biol. 2013. PMID: 23748610 Free PMC article.
-
Heterochromatin Protein 1β (HP1β) has distinct functions and distinct nuclear distribution in pluripotent versus differentiated cells.Genome Biol. 2015 Sep 28;16:213. doi: 10.1186/s13059-015-0760-8. Genome Biol. 2015. PMID: 26415775 Free PMC article.
-
HP1γ regulates H3K36 methylation and pluripotency in embryonic stem cells.Nucleic Acids Res. 2020 Dec 16;48(22):12660-12674. doi: 10.1093/nar/gkaa1091. Nucleic Acids Res. 2020. PMID: 33237287 Free PMC article.
-
Diverse roles of WDR5-RbBP5-ASH2L-DPY30 (WRAD) complex in the functions of the SET1 histone methyltransferase family.J Biosci. 2017 Mar;42(1):155-159. doi: 10.1007/s12038-017-9666-9. J Biosci. 2017. PMID: 28229975 Review.
-
Function of heterochromatin protein 1 during DNA repair.Protoplasma. 2017 May;254(3):1233-1240. doi: 10.1007/s00709-017-1090-3. Epub 2017 Feb 24. Protoplasma. 2017. PMID: 28236007 Review.
Cited by
-
DPPA2, DPPA4, and other DPPA factor epigenomic functions in cell fate and cancer.Stem Cell Reports. 2021 Dec 14;16(12):2844-2851. doi: 10.1016/j.stemcr.2021.10.008. Epub 2021 Nov 11. Stem Cell Reports. 2021. PMID: 34767751 Free PMC article. Review.
-
KDM3A and KDM3B Maintain Naïve Pluripotency Through the Regulation of Alternative Splicing.bioRxiv [Preprint]. 2024 Jan 23:2023.05.31.543088. doi: 10.1101/2023.05.31.543088. bioRxiv. 2024. PMID: 37398291 Free PMC article. Preprint.
-
N6-methyladenosine-modified SENP1, identified by IGF2BP3, is a novel molecular marker in acute myeloid leukemia and aggravates progression by activating AKT signal via de-SUMOylating HDAC2.Mol Cancer. 2024 May 31;23(1):116. doi: 10.1186/s12943-024-02013-y. Mol Cancer. 2024. PMID: 38822351 Free PMC article.
-
Heterochromatin protein 1 beta regulates neural and neural crest development by repressing pluripotency-associated gene pou5f3.2/oct25 in Xenopus.Dev Dyn. 2021 Aug;250(8):1113-1124. doi: 10.1002/dvdy.319. Epub 2021 Feb 26. Dev Dyn. 2021. PMID: 33595886 Free PMC article.
-
A balanced Oct4 interactome is crucial for maintaining pluripotency.Sci Adv. 2022 Feb 18;8(7):eabe4375. doi: 10.1126/sciadv.abe4375. Epub 2022 Feb 16. Sci Adv. 2022. PMID: 35171666 Free PMC article.
References
-
- Ahmad K., Henikoff S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell. 2002;9:1191–1200. - PubMed
-
- Bannister A., Zegerman P., Partridge J., Miska E., Thomas J., Allshire R. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature. 2001;410:120–124. - PubMed
-
- Beard C., Hochedlinger K., Plath K., Wutz A., Jaenisch R. Efficient method to generate single-copy transgenic mice by site-specific integration in embryonic stem cells. Genesis. 2006;44:23–28. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

