History of CRISPR-Cas from Encounter with a Mysterious Repeated Sequence to Genome Editing Technology
- PMID: 29358495
- PMCID: PMC5847661
- DOI: 10.1128/JB.00580-17
History of CRISPR-Cas from Encounter with a Mysterious Repeated Sequence to Genome Editing Technology
Abstract
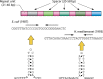
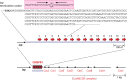
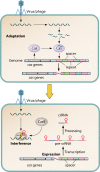
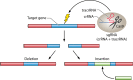
Clustered regularly interspaced short palindromic repeat (CRISPR)-Cas systems are well-known acquired immunity systems that are widespread in archaea and bacteria. The RNA-guided nucleases from CRISPR-Cas systems are currently regarded as the most reliable tools for genome editing and engineering. The first hint of their existence came in 1987, when an unusual repetitive DNA sequence, which subsequently was defined as a CRISPR, was discovered in the Escherichia coli genome during an analysis of genes involved in phosphate metabolism. Similar sequence patterns were then reported in a range of other bacteria as well as in halophilic archaea, suggesting an important role for such evolutionarily conserved clusters of repeated sequences. A critical step toward functional characterization of the CRISPR-Cas systems was the recognition of a link between CRISPRs and the associated Cas proteins, which were initially hypothesized to be involved in DNA repair in hyperthermophilic archaea. Comparative genomics, structural biology, and advanced biochemistry could then work hand in hand, not only culminating in the explosion of genome editing tools based on CRISPR-Cas9 and other class II CRISPR-Cas systems but also providing insights into the origin and evolution of this system from mobile genetic elements denoted casposons. To celebrate the 30th anniversary of the discovery of CRISPR, this minireview briefly discusses the fascinating history of CRISPR-Cas systems, from the original observation of an enigmatic sequence in E. coli to genome editing in humans.
Keywords: archaea; casposon; genome editing; repeated sequence.
Copyright © 2018 American Society for Microbiology.
Figures


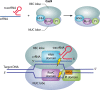



References
-
- Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. 1987. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol 169:5429–5433. doi: 10.1128/jb.169.12.5429-5433.1987. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

