Genetic Analysis of Serum-Derived Defective Hepatitis C Virus Genomes Revealed Novel Viral cis Elements for Virus Replication and Assembly
- PMID: 29367245
- PMCID: PMC5972877
- DOI: 10.1128/JVI.02182-17
Genetic Analysis of Serum-Derived Defective Hepatitis C Virus Genomes Revealed Novel Viral cis Elements for Virus Replication and Assembly
Abstract
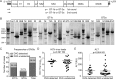
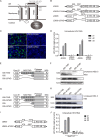
Defective viral genomes (DVGs) of hepatitis C virus (HCV) exist, but their biological significances have not been thoroughly investigated. Here, we analyzed HCV DVGs circulating in patient sera that possess deletions in the structural protein-encoding region. About 30% of 41 HCV clinical isolates possess DVGs that originated from the full-length genome in the same patients. No correlation between DVGs, viremia, and alanine aminotransferase (ALT) levels was found. Sequencing analysis of DVGs revealed the existence of deletion hot spots, with upstream sites in E1 and downstream sites in E2 and NS2. Interestingly, the coding sequences for the core protein and the C-terminal protease domain of NS2 were always intact in DVGs despite the fact that both proteins are dispensable for HCV genome replication. Mechanistic studies showed that transmembrane segment 3 (TMS3) of NS2, located immediately upstream of its protease domain, was required for the cleavage of NS2-NS3 and the replication of DVGs. Moreover, we identified a highly conserved secondary structure (SL750) within the core domain 2-coding region that is critical for HCV genome packaging. In summary, our analysis of serum-derived HCV DVGs revealed novel viral cis elements that play important roles in virus replication and assembly.IMPORTANCE HCV DVGs have been identified in vivo and in vitro, but their biogenesis and physiological significances remain elusive. In addition, a conventional packaging signal has not yet been identified on the HCV RNA genome, and mechanisms underlying the specificity in the encapsidation of the HCV genome into infectious particles remain to be uncovered. Here, we identified new viral cis elements critical for the HCV life cycle by determining genetic constraints that define the boundary of serum-derived HCV DVGs. We found that transmembrane segment 3 of NS2, located immediately upstream of its protease domain, was required for the cleavage of NS2-NS3 and the replication of DVGs. We identified a highly conserved secondary structure (SL750) within the core-coding region that is critical for HCV genome packaging. In summary, our analysis of serum-derived HCV DVGs revealed previously unexpected novel cis elements critical for HCV replication and morphogenesis.
Keywords: NS2; cis elements; core; defective genome; hepatitis C virus; virus packaging.
Copyright © 2018 American Society for Microbiology.
Figures






References
-
- Li D, Huang Z, Zhong J. 2015. Hepatitis C virus vaccine development: old challenges and new opportunities. Nat Sci Rev 2:285–295. doi: 10.1093/nsr/nwv040. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

