Plastid phylogenomics with broad taxon sampling further elucidates the distinct evolutionary origins and timing of secondary green plastids
- PMID: 29367699
- PMCID: PMC5784168
- DOI: 10.1038/s41598-017-18805-w
Plastid phylogenomics with broad taxon sampling further elucidates the distinct evolutionary origins and timing of secondary green plastids
Abstract
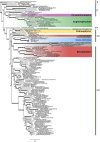
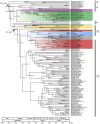
Secondary plastids derived from green algae occur in chlorarachniophytes, photosynthetic euglenophytes, and the dinoflagellate genus Lepidodinium. Recent advances in understanding the origin of these plastids have been made, but analyses suffer from relatively sparse taxon sampling within the green algal groups to which they are related. In this study we aim to derive new insights into the identity of the plastid donors, and when in geological time the independent endosymbiosis events occurred. We use newly sequenced green algal chloroplast genomes from carefully chosen lineages potentially related to chlorarachniophyte and Lepidodinium plastids, combined with recently published chloroplast genomes, to present taxon-rich phylogenetic analyses to further pinpoint plastid origins. We integrate phylogenies with fossil information and relaxed molecular clock analyses. Our results indicate that the chlorarachniophyte plastid may originate from a precusor of siphonous green algae or a closely related lineage, whereas the Lepidodinium plastid originated from a pedinophyte. The euglenophyte plastid putatively originated from a lineage of prasinophytes within the order Pyramimonadales. Our molecular clock analyses narrow in on the likely timing of the secondary endosymbiosis events, suggesting that the event leading to Lepidodinium likely occurred more recently than those leading to the chlorarachniophyte and photosynthetic euglenophyte lineages.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

