Widespread signatures of selection for secreted peptidases in a fungal plant pathogen
- PMID: 29368587
- PMCID: PMC5784588
- DOI: 10.1186/s12862-018-1123-3
Widespread signatures of selection for secreted peptidases in a fungal plant pathogen
Abstract
Background: Fungal plant pathogens secrete a large arsenal of hydrolytic enzymes during the course of infection, including peptidases. Secreted peptidases have been extensively studied for their role as effectors. In this study, we combined transcriptomics, comparative genomics and evolutionary analyses to investigate all 39 secreted peptidases in the fungal wheat pathogen Zymoseptoria tritici and its close relatives Z. pseudotritici and Z. ardabiliae.
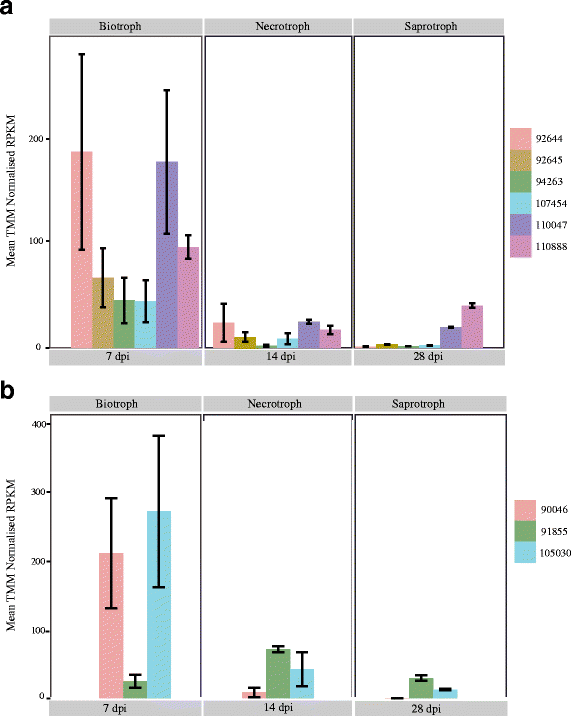
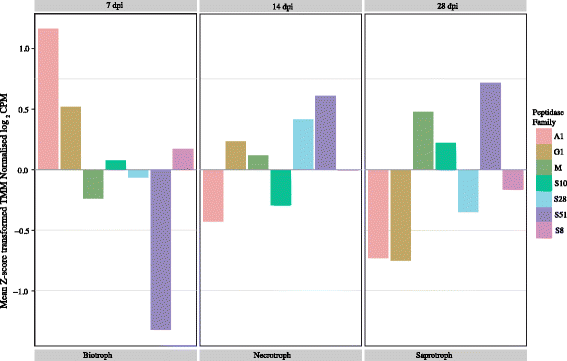
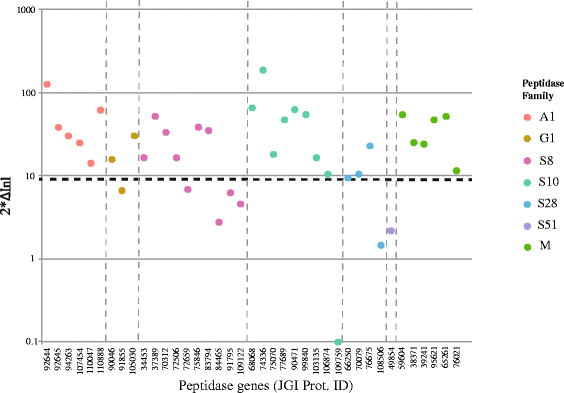
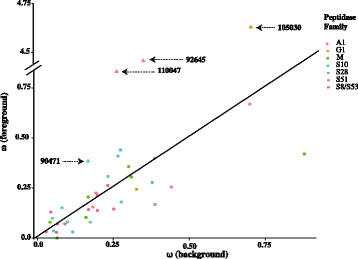
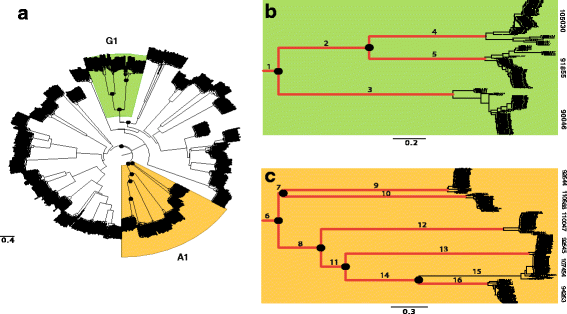
Results: RNA-seq data revealed that a majority of the secreted peptidases displayed differential transcription during the course of Z. tritici infection, indicative of specialization for different stages in the life cycle. Evolutionary analyses detected widespread evidence of adaptive evolution acting on at least 28 of the peptidases. A few peptidases displayed lineage-specific rates of molecular evolution, suggesting altered selection pressure in Z. tritici following host specialization on domesticated wheat. The peptidases belonging to MEROPS families A1 and G1 emerged as a particularly interesting group that may play key roles in host-pathogen co-evolution, host adaptation and pathogenicity. Sister genes in the A1 and G1 families showed accelerated substitution rates after gene duplications.
Conclusions: These results suggest widespread evolution of secreted peptidases leading to novel gene functions, consistent with predicted models of "escape from adaptive conflict" and "neo-functionalization". Our analyses identified candidate genes worthy of functional analyses that may encode effector functions, for example by suppressing plant defenses during the biotrophic phase of infection.
Keywords: Adaptive evolution; Escape from adaptive conflict (EAC); Neo-functionalization (NEO-F); Peptidases; Transcriptomics.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
Similar articles
-
Coevolution and life cycle specialization of plant cell wall degrading enzymes in a hemibiotrophic pathogen.Mol Biol Evol. 2013 Jun;30(6):1337-47. doi: 10.1093/molbev/mst041. Epub 2013 Mar 20. Mol Biol Evol. 2013. PMID: 23515261 Free PMC article.
-
Rapidly Evolving Genes Are Key Players in Host Specialization and Virulence of the Fungal Wheat Pathogen Zymoseptoria tritici (Mycosphaerella graminicola).PLoS Pathog. 2015 Jul 30;11(7):e1005055. doi: 10.1371/journal.ppat.1005055. eCollection 2015 Jul. PLoS Pathog. 2015. PMID: 26225424 Free PMC article.
-
Expression profiling of the wheat pathogen Zymoseptoria tritici reveals genomic patterns of transcription and host-specific regulatory programs.Genome Biol Evol. 2014 May 14;6(6):1353-65. doi: 10.1093/gbe/evu101. Genome Biol Evol. 2014. PMID: 24920004 Free PMC article.
-
How Knowledge of Pathogen Population Biology Informs Management of Septoria Tritici Blotch.Phytopathology. 2016 Sep;106(9):948-55. doi: 10.1094/PHYTO-03-16-0131-RVW. Epub 2016 Jul 27. Phytopathology. 2016. PMID: 27111799 Review.
-
Stagonospora nodorum: from pathology to genomics and host resistance.Annu Rev Phytopathol. 2012;50:23-43. doi: 10.1146/annurev-phyto-081211-173019. Epub 2012 May 1. Annu Rev Phytopathol. 2012. PMID: 22559071 Review.
Cited by
-
Defense and Offense Strategies: The Role of Aspartic Proteases in Plant-Pathogen Interactions.Biology (Basel). 2021 Jan 21;10(2):75. doi: 10.3390/biology10020075. Biology (Basel). 2021. PMID: 33494266 Free PMC article. Review.
-
Necrotrophic lifestyle of Rhizoctonia solani AG3-PT during interaction with its host plant potato as revealed by transcriptome analysis.Sci Rep. 2020 Jul 28;10(1):12574. doi: 10.1038/s41598-020-68728-2. Sci Rep. 2020. PMID: 32724205 Free PMC article.
-
In silico analysis of secreted effectorome of the rubber tree pathogen Rigidoporus microporus highlights its potential virulence proteins.Front Microbiol. 2024 Sep 16;15:1439454. doi: 10.3389/fmicb.2024.1439454. eCollection 2024. Front Microbiol. 2024. PMID: 39360316 Free PMC article.
-
Large-scale genomic analyses with machine learning uncover predictive patterns associated with fungal phytopathogenic lifestyles and traits.Sci Rep. 2023 Oct 11;13(1):17203. doi: 10.1038/s41598-023-44005-w. Sci Rep. 2023. PMID: 37821494 Free PMC article.
-
Genome-Wide Association Study Reveals Novel Powdery Mildew Resistance Loci in Bread Wheat.Plants (Basel). 2023 Nov 15;12(22):3864. doi: 10.3390/plants12223864. Plants (Basel). 2023. PMID: 38005757 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

