2-Phenylquinazolinones as dual-activity tankyrase-kinase inhibitors
- PMID: 29374194
- PMCID: PMC5785997
- DOI: 10.1038/s41598-018-19872-3
2-Phenylquinazolinones as dual-activity tankyrase-kinase inhibitors
Abstract
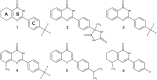
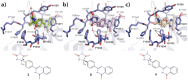
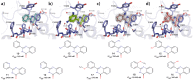
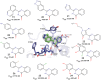
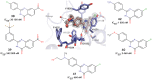
Tankyrases (TNKSs) are enzymes specialized in catalyzing poly-ADP-ribosylation of target proteins. Several studies have validated TNKSs as anti-cancer drug targets due to their regulatory role in Wnt/β-catenin pathway. Recently a lot of effort has been put into developing more potent and selective TNKS inhibitors and optimizing them towards anti-cancer agents. We noticed that some 2-phenylquinazolinones (2-PQs) reported as CDK9 inhibitors were similar to previously published TNKS inhibitors. In this study, we profiled this series of 2-PQs against TNKS and selected kinases that are involved in the Wnt/β-catenin pathway. We found that they were much more potent TNKS inhibitors than they were CDK9/kinase inhibitors. We evaluated the compound selectivity to tankyrases over the ARTD enzyme family and solved co-crystal structures of the compounds with TNKS2. Comparative structure-based studies of the catalytic domain of TNKS2 with selected CDK9 inhibitors and docking studies of the inhibitors with two kinases (CDK9 and Akt) revealed important structural features, which could explain the selectivity of the compounds towards either tankyrases or kinases. We also discovered a compound, which was able to inhibit tankyrases, CDK9 and Akt kinases with equal µM potency.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


Similar articles
-
Therapeutic Path to Triple Knockout: Investigating the Pan-inhibitory Mechanisms of AKT, CDK9, and TNKS2 by a Novel 2-phenylquinazolinone Derivative in Cancer Therapy- An In-silico Investigation Therapy.Curr Pharm Biotechnol. 2024;25(10):1288-1303. doi: 10.2174/1389201024666230815145001. Curr Pharm Biotechnol. 2024. PMID: 37581526
-
Structure-activity relationships of 2-arylquinazolin-4-ones as highly selective and potent inhibitors of the tankyrases.Eur J Med Chem. 2016 Aug 8;118:316-27. doi: 10.1016/j.ejmech.2016.04.041. Epub 2016 Apr 20. Eur J Med Chem. 2016. PMID: 27163581
-
Molecular insights on TNKS1/TNKS2 and inhibitor-IWR1 interactions.Mol Biosyst. 2014 Feb;10(2):281-93. doi: 10.1039/c3mb70305c. Mol Biosyst. 2014. PMID: 24291818
-
Insights of tankyrases: A novel target for drug discovery.Eur J Med Chem. 2020 Dec 1;207:112712. doi: 10.1016/j.ejmech.2020.112712. Epub 2020 Aug 17. Eur J Med Chem. 2020. PMID: 32877803 Review.
-
Tankyrases as drug targets.FEBS J. 2013 Aug;280(15):3576-93. doi: 10.1111/febs.12320. Epub 2013 Jun 18. FEBS J. 2013. PMID: 23648170 Review.
Cited by
-
3,4,3'-Tri-O-methylellagic acid as an anticancer agent: in vitro and in silico studies.RSC Adv. 2022 Oct 19;12(46):29884-29891. doi: 10.1039/d2ra05246f. eCollection 2022 Oct 17. RSC Adv. 2022. PMID: 36321100 Free PMC article.
-
Structural basis of tankyrase activation by polymerization.Nature. 2022 Dec;612(7938):162-169. doi: 10.1038/s41586-022-05449-8. Epub 2022 Nov 23. Nature. 2022. PMID: 36418402 Free PMC article.
-
[1,2,4]Triazolo[3,4-b]benzothiazole Scaffold as Versatile Nicotinamide Mimic Allowing Nanomolar Inhibition of Different PARP Enzymes.J Med Chem. 2023 Jan 26;66(2):1301-1320. doi: 10.1021/acs.jmedchem.2c01460. Epub 2023 Jan 4. J Med Chem. 2023. PMID: 36598465 Free PMC article.
-
Exploring the Effects of Chirality of 5-methyl-5-[4-(4-oxo-3H-quinazolin-2- yl)phenyl]imidazolidine-2,4-dione and its Derivatives on the Oncological Target Tankyrase 2. Atomistic Insights.Curr Pharm Biotechnol. 2025;26(2):222-234. doi: 10.2174/1389201024666230330084017. Curr Pharm Biotechnol. 2025. PMID: 37005548
-
Discovery of Novel Inhibitor for WNT/β-Catenin Pathway by Tankyrase 1/2 Structure-Based Virtual Screening.Molecules. 2020 Apr 6;25(7):1680. doi: 10.3390/molecules25071680. Molecules. 2020. PMID: 32268564 Free PMC article.
References
-
- Matsutani N, et al. Expression of telomeric repeat binding factor 1 and 2 and TRF1-interacting nuclear protein 2 in human gastric carcinomas. Int. J. Oncol. 2001;19:507–512. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

