Quantitative or qualitative transcriptional diagnostic signatures? A case study for colorectal cancer
- PMID: 29378509
- PMCID: PMC5789529
- DOI: 10.1186/s12864-018-4446-y
Quantitative or qualitative transcriptional diagnostic signatures? A case study for colorectal cancer
Abstract
Background: Due to experimental batch effects, the application of a quantitative transcriptional signature for disease diagnoses commonly requires inter-sample data normalization, which would be hardly applicable under common clinical settings. Many cancers might have qualitative differences with the non-cancer states in the gene expression pattern. Therefore, it is reasonable to explore the power of qualitative diagnostic signatures which are robust against experimental batch effects and other random factors.
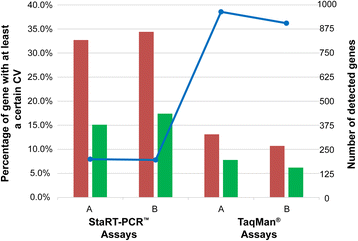
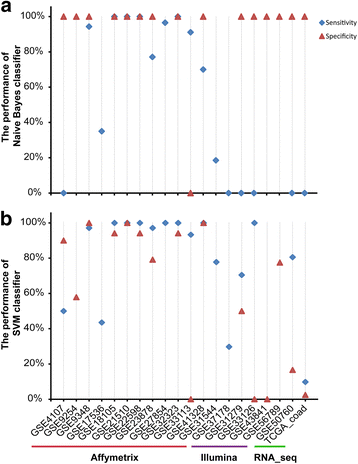
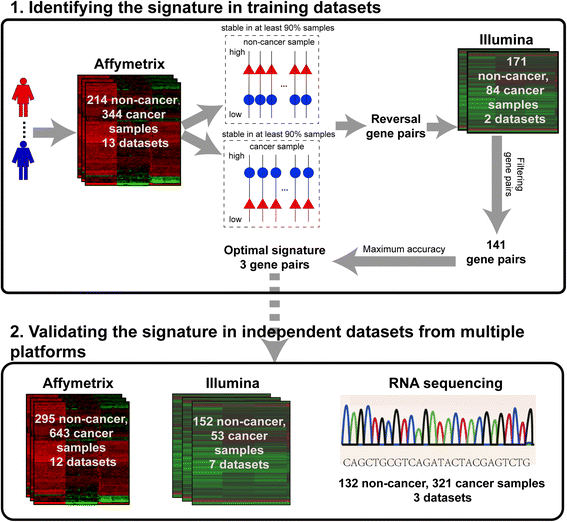
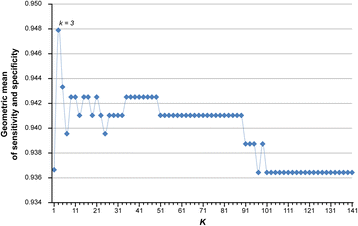
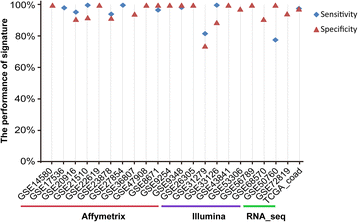
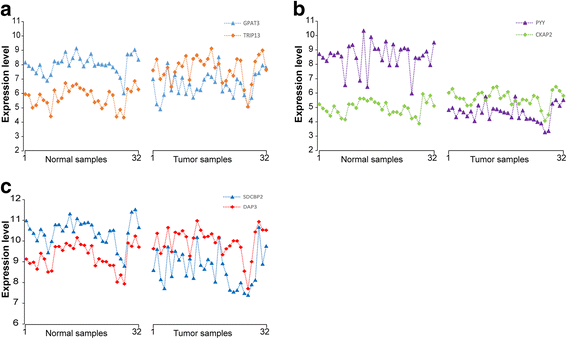
Results: Firstly, using data of technical replicate samples from the MicroArray Quality Control (MAQC) project, we demonstrated that the low-throughput PCR-based technologies also exist large measurement variations for gene expression even when the samples were measured in the same test site. Then, we demonstrated the critical limitation of low stability for classifiers based on quantitative transcriptional signatures in applications to individual samples through a case study using a support vector machine and a naïve Bayesian classifier to discriminate colorectal cancer tissues from normal tissues. To address this problem, we identified a signature consisting of three gene pairs for discriminating colorectal cancer tissues from non-cancer (normal and inflammatory bowel disease) tissues based on within-sample relative expression orderings (REOs) of these gene pairs. The signature was well verified using 22 independent datasets measured by different microarray and RNA_seq platforms, obviating the need of inter-sample data normalization.
Conclusions: Subtle quantitative information of gene expression measurements tends to be unstable under current technical conditions, which will introduce uncertainty to clinical applications of the quantitative transcriptional diagnostic signatures. For diagnosis of disease states with qualitative transcriptional characteristics, the qualitative REO-based signatures could be robustly applied to individual samples measured by different platforms.
Keywords: Batch effects; Classifiers; Diagnostic signature; Platform; Relative expression orderings.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no conflict of interest.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures
Similar articles
-
Two novel qualitative transcriptional signatures robustly applicable to non-research-oriented colorectal cancer samples with low-quality RNA.J Cell Mol Med. 2021 Apr;25(7):3622-3633. doi: 10.1111/jcmm.16467. Epub 2021 Mar 14. J Cell Mol Med. 2021. PMID: 33719152 Free PMC article.
-
A qualitative transcriptional signature for the early diagnosis of colorectal cancer.Cancer Sci. 2019 Oct;110(10):3225-3234. doi: 10.1111/cas.14137. Epub 2019 Sep 3. Cancer Sci. 2019. PMID: 31335996 Free PMC article.
-
Circumvent the uncertainty in the applications of transcriptional signatures to tumor tissues sampled from different tumor sites.Oncotarget. 2017 May 2;8(18):30265-30275. doi: 10.18632/oncotarget.15754. Oncotarget. 2017. PMID: 28427173 Free PMC article.
-
Why breast cancer signatures are no better than random signatures explained.Drug Discov Today. 2018 Nov;23(11):1818-1823. doi: 10.1016/j.drudis.2018.05.036. Epub 2018 Jun 1. Drug Discov Today. 2018. PMID: 29864526 Review.
-
Molecular subtyping of cancer: current status and moving toward clinical applications.Brief Bioinform. 2019 Mar 25;20(2):572-584. doi: 10.1093/bib/bby026. Brief Bioinform. 2019. PMID: 29659698 Review.
Cited by
-
Individualized detection of TMPRSS2-ERG fusion status in prostate cancer: a rank-based qualitative transcriptome signature.World J Surg Oncol. 2024 Feb 9;22(1):49. doi: 10.1186/s12957-024-03314-8. World J Surg Oncol. 2024. PMID: 38331878 Free PMC article.
-
Emergence of Cardiac Glycosides as Potential Drugs: Current and Future Scope for Cancer Therapeutics.Biomolecules. 2021 Aug 25;11(9):1275. doi: 10.3390/biom11091275. Biomolecules. 2021. PMID: 34572488 Free PMC article. Review.
-
Qualitative transcriptional signature for predicting pathological response of colorectal cancer to FOLFOX therapy.Cancer Sci. 2020 Jan;111(1):253-265. doi: 10.1111/cas.14263. Epub 2019 Dec 18. Cancer Sci. 2020. PMID: 31785020 Free PMC article.
-
A qualitative transcriptional signature for the histological reclassification of lung squamous cell carcinomas and adenocarcinomas.BMC Genomics. 2019 Nov 21;20(1):881. doi: 10.1186/s12864-019-6086-2. BMC Genomics. 2019. PMID: 31752667 Free PMC article.
-
Hierarchical identification of a transcriptional panel for the histological diagnosis of lung neuroendocrine tumors.Front Genet. 2022 Aug 29;13:944167. doi: 10.3389/fgene.2022.944167. eCollection 2022. Front Genet. 2022. PMID: 36105102 Free PMC article.
References
-
- Brawley OW, Flenaugh EL. Low-dose spiral CT screening and evaluation of the solitary pulmonary nodule. Oncology (Williston Park) 2014;28(5):441–446. - PubMed
-
- Wang YR, Cangemi JR, Loftus EV, Jr., Picco MF: Rate of early/missed colorectal cancers after colonoscopy in older patients with or without inflammatory bowel disease in the United States. Am J Gastroenterol 2013, 108(3):444–449. - PubMed
-
- European Colorectal Cancer Screening Guidelines Working Group. von Karsa L, Patnick J, Segnan N, Atkin W, Halloran S, Lansdorp-Vogelaar I, Malila N, Minozzi S, Moss S, et al. European guidelines for quality assurance in colorectal cancer screening and diagnosis: overview and introduction to the full supplement publication. Endoscopy. 2013;45(1):51–59. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 81372213/National Natural Science Foundation of China/International
- 81572935/National Natural Science Foundation of China/International
- 21534008/National Natural Science Foundation of China/International
- 61601151/National Natural Science Foundation of China/International
- 81602738/National Natural Science Foundation of China (CN)/International
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

