Differential depth distribution of microbial function and putative symbionts through sediment-hosted aquifers in the deep terrestrial subsurface
- PMID: 29379208
- PMCID: PMC6792436
- DOI: 10.1038/s41564-017-0098-y
Differential depth distribution of microbial function and putative symbionts through sediment-hosted aquifers in the deep terrestrial subsurface
Abstract
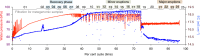
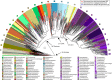
An enormous diversity of previously unknown bacteria and archaea has been discovered recently, yet their functional capacities and distributions in the terrestrial subsurface remain uncertain. Here, we continually sampled a CO2-driven geyser (Colorado Plateau, Utah, USA) over its 5-day eruption cycle to test the hypothesis that stratified, sandstone-hosted aquifers sampled over three phases of the eruption cycle have microbial communities that differ both in membership and function. Genome-resolved metagenomics, single-cell genomics and geochemical analyses confirmed this hypothesis and linked microorganisms to groundwater compositions from different depths. Autotrophic Candidatus "Altiarchaeum sp." and phylogenetically deep-branching nanoarchaea dominate the deepest groundwater. A nanoarchaeon with limited metabolic capacity is inferred to be a potential symbiont of the Ca. "Altiarchaeum". Candidate Phyla Radiation bacteria are also present in the deepest groundwater and they are relatively abundant in water from intermediate depths. During the recovery phase of the geyser, microaerophilic Fe- and S-oxidizers have high in situ genome replication rates. Autotrophic Sulfurimonas sustained by aerobic sulfide oxidation and with the capacity for N2 fixation dominate the shallow aquifer. Overall, 104 different phylum-level lineages are present in water from these subsurface environments, with uncultivated archaea and bacteria partitioned to the deeper subsurface.
Conflict of interest statement
The authors declare no competing financial interests.
Figures




References
-
- Pedersen K, Arlinger J, Ekendahl S, Hallbeck L. 16S rRNA gene diversity of attached and unattached bacteria in boreholes along the access tunnel to the Äspö hard rock laboratory, Sweden. Fems. Microbiol. Ecol. 1996;19:249–262.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

