The SR protein B52/SRp55 regulates splicing of the period thermosensitive intron and mid-day siesta in Drosophila
- PMID: 29382842
- PMCID: PMC5789894
- DOI: 10.1038/s41598-017-18167-3
The SR protein B52/SRp55 regulates splicing of the period thermosensitive intron and mid-day siesta in Drosophila
Abstract
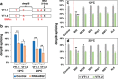
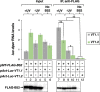
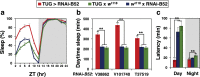
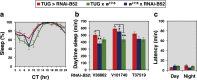
Similar to many diurnal animals, Drosophila melanogaster exhibits a mid-day siesta that is more robust as temperature increases, an adaptive response that aims to minimize the deleterious effects from exposure to heat. This temperature-dependent plasticity in mid-day sleep levels is partly based on the thermal sensitive splicing of an intron in the 3' untranslated region (UTR) of the circadian clock gene termed period (per). In this study, we evaluated a possible role for the serine/arginine-rich (SR) splicing factors in the regulation of dmpi8 splicing efficiency and mid-day siesta. Using a Drosophila cell culture assay we show that B52/SRp55 increases dmpi8 splicing efficiency, whereas other SR proteins have little to no effect. The magnitude of the stimulatory effect of B52 on dmpi8 splicing efficiency is modulated by natural variation in single nucleotide polymorphisms (SNPs) in the per 3' UTR that correlate with B52 binding levels. Down-regulating B52 expression in clock neurons increases mid-day siesta and reduces dmpi8 splicing efficiency. Our results establish a novel role for SR proteins in sleep and suggest that polymorphisms in the per 3' UTR contribute to natural variation in sleep behavior by modulating the binding efficiencies of SR proteins.
Conflict of interest statement
The authors declare that they have no competing interests.
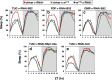
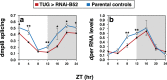
Figures
Similar articles
-
Natural variation in the Drosophila melanogaster clock gene period modulates splicing of its 3'-terminal intron and mid-day siesta.PLoS One. 2012;7(11):e49536. doi: 10.1371/journal.pone.0049536. Epub 2012 Nov 13. PLoS One. 2012. PMID: 23152918 Free PMC article.
-
Mid-day siesta in natural populations of D. melanogaster from Africa exhibits an altitudinal cline and is regulated by splicing of a thermosensitive intron in the period clock gene.BMC Evol Biol. 2017 Jan 23;17(1):32. doi: 10.1186/s12862-017-0880-8. BMC Evol Biol. 2017. PMID: 28114910 Free PMC article.
-
Parallel clinal variation in the mid-day siesta of Drosophila melanogaster implicates continent-specific targets of natural selection.PLoS Genet. 2018 Sep 4;14(9):e1007612. doi: 10.1371/journal.pgen.1007612. eCollection 2018 Sep. PLoS Genet. 2018. PMID: 30180162 Free PMC article.
-
Thermosensitive splicing of a clock gene and seasonal adaptation.Cold Spring Harb Symp Quant Biol. 2007;72:599-606. doi: 10.1101/sqb.2007.72.021. Cold Spring Harb Symp Quant Biol. 2007. PMID: 18419319 Review.
-
Circadian Rhythms and Sleep in Drosophila melanogaster.Genetics. 2017 Apr;205(4):1373-1397. doi: 10.1534/genetics.115.185157. Genetics. 2017. PMID: 28360128 Free PMC article. Review.
Cited by
-
Sleep in Drosophila and Its Context.Front Physiol. 2019 Sep 11;10:1167. doi: 10.3389/fphys.2019.01167. eCollection 2019. Front Physiol. 2019. PMID: 31572216 Free PMC article. Review.
-
Integration of photoperiodic and temperature cues by the circadian clock to regulate insect seasonal adaptations.J Comp Physiol A Neuroethol Sens Neural Behav Physiol. 2024 Jul;210(4):585-599. doi: 10.1007/s00359-023-01667-1. Epub 2023 Aug 16. J Comp Physiol A Neuroethol Sens Neural Behav Physiol. 2024. PMID: 37584703 Free PMC article. Review.
-
Post-transcriptional modulators and mediators of the circadian clock.Chronobiol Int. 2021 Sep;38(9):1244-1261. doi: 10.1080/07420528.2021.1928159. Epub 2021 May 31. Chronobiol Int. 2021. PMID: 34056966 Free PMC article. Review.
-
Drosophila PSI controls circadian period and the phase of circadian behavior under temperature cycle via tim splicing.Elife. 2019 Nov 8;8:e50063. doi: 10.7554/eLife.50063. Elife. 2019. PMID: 31702555 Free PMC article.
-
Daywake, an Anti-siesta Gene Linked to a Splicing-Based Thermostat from an Adjoining Clock Gene.Curr Biol. 2019 May 20;29(10):1728-1734.e4. doi: 10.1016/j.cub.2019.04.039. Epub 2019 May 9. Curr Biol. 2019. PMID: 31080079 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

