Identification of key candidate genes and pathways in hepatitis B virus-associated acute liver failure by bioinformatical analysis
- PMID: 29384847
- PMCID: PMC5805419
- DOI: 10.1097/MD.0000000000009687
Identification of key candidate genes and pathways in hepatitis B virus-associated acute liver failure by bioinformatical analysis
Abstract
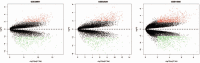
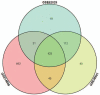
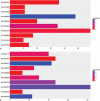
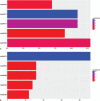
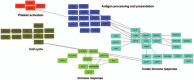
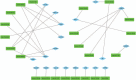
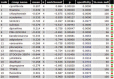
Hepatitis B virus-associated acute liver failure (HBV-ALF) is a rare but life-threatening syndrome that carried a high morbidity and mortality. Our study aimed to explore the possible molecular mechanisms of HBV-ALF by means of bioinformatics analysis. In this study, genes expression microarray datasets of HBV-ALF from Gene Expression Omnibus were collected, and then we identified differentially expressed genes (DEGs) by the limma package in R. After functional enrichment analysis, we constructed the protein-protein interaction (PPI) network by the Search Tool for the Retrieval of Interacting Genes online database and weighted genes coexpression network by the WGCNA package in R. Subsequently, we picked out the hub genes among the DEGs. A total of 423 DEGs with 198 upregulated genes and 225 downregulated genes were identified between HBV-ALF and normal samples. The upregulated genes were mainly enriched in immune response, and the downregulated genes were mainly enriched in complement and coagulation cascades. Orosomucoid 1 (ORM1), orosomucoid 2 (ORM2), plasminogen (PLG), and aldehyde oxidase 1 (AOX1) were picked out as the hub genes that with a high degree in both PPI network and weighted genes coexpression network. The weighted genes coexpression network analysis found out 3 of the 5 modules that upregulated genes enriched in were closely related to immune system. The downregulated genes enriched in only one module, and the genes in this module majorly enriched in the complement and coagulation cascades pathway. In conclusion, 4 genes (ORM1, ORM2, PLG, and AOX1) with immune response and the complement and coagulation cascades pathway may take part in the pathogenesis of HBV-ALF, and these candidate genes and pathways could be therapeutic targets for HBV-ALF.
Conflict of interest statement
The authors have no conflicts of interest to disclose.
Figures



Similar articles
-
Potential Biomarkers and Therapeutic Targets in Hepatitis B Virus-related Acute Liver Failure: Interplay of the Ferroptosis, Autophagy and Immune Responses.Int J Med Sci. 2025 Jan 21;22(4):806-818. doi: 10.7150/ijms.106360. eCollection 2025. Int J Med Sci. 2025. PMID: 39991755 Free PMC article.
-
Systematic analysis on multiple Gene Expression Omnibus data sets reveals fierce immune response in hepatitis B virus-related acute liver failure.J Cell Mol Med. 2020 Sep;24(17):9798-9809. doi: 10.1111/jcmm.15561. Epub 2020 Jul 19. J Cell Mol Med. 2020. PMID: 32686296 Free PMC article.
-
Bioinformatic analysis of differentially expressed genes involved in the hepatitis B virus-associated acute liver failure.Acta Gastroenterol Belg. 2018 Apr-Jun;81(2):288-294. Acta Gastroenterol Belg. 2018. PMID: 30024701
-
The regulatory role of microRNA-mRNA co-expression in hepatitis B virus-associated acute liver failure.Ann Hepatol. 2019 Nov-Dec;18(6):883-892. doi: 10.1016/j.aohep.2019.07.007. Epub 2019 Aug 21. Ann Hepatol. 2019. PMID: 31521462
-
Bioinformatics and experimental validation were combined to explore lactylation-related biomarkers in HBV-associated acute liver failure.J Gastroenterol Hepatol. 2024 Dec;39(12):2903-2915. doi: 10.1111/jgh.16739. Epub 2024 Sep 16. J Gastroenterol Hepatol. 2024. PMID: 39285310
Cited by
-
Bioinformatics analysis on multiple Gene Expression Omnibus datasets of the hepatitis B virus infection and its response to the interferon-alpha therapy.BMC Infect Dis. 2020 Jan 29;20(1):84. doi: 10.1186/s12879-019-4720-x. BMC Infect Dis. 2020. PMID: 31996147 Free PMC article.
-
High glycosylated serum protein to high density lipoprotein cholesterol ratios are predictive of worse acute on chronic liver failure prognoses.Sci Rep. 2025 Apr 24;15(1):14288. doi: 10.1038/s41598-025-91779-2. Sci Rep. 2025. PMID: 40274908 Free PMC article.
-
Potential Networks Regulated by MSCs in Acute-On-Chronic Liver Failure: Exosomal miRNAs and Intracellular Target Genes.Front Genet. 2021 Apr 23;12:650536. doi: 10.3389/fgene.2021.650536. eCollection 2021. Front Genet. 2021. PMID: 33968135 Free PMC article.
-
Complement System as a Target for Therapies to Control Liver Regeneration/Damage in Acute Liver Failure Induced by Viral Hepatitis.J Immunol Res. 2018 Oct 8;2018:3917032. doi: 10.1155/2018/3917032. eCollection 2018. J Immunol Res. 2018. PMID: 30402508 Free PMC article.
-
Downregulation of orosomucoid 2 acts as a prognostic factor associated with cancer-promoting pathways in liver cancer.World J Gastroenterol. 2020 Feb 28;26(8):804-817. doi: 10.3748/wjg.v26.i8.804. World J Gastroenterol. 2020. PMID: 32148378 Free PMC article.
References
-
- Liu J, Fan D. Hepatitis B in China. Lancet 2007;369:1582–3. - PubMed
-
- Escorsell A, Mas A, de la Mata M. Acute liver failure in Spain: analysis of 267 cases. Liver Transpl 2007;13:1389–95. - PubMed
-
- Koskinas J, Deutsch M, Kountouras D, et al. Aetiology and outcome of acute hepatic failure in Greece: experience of two academic hospital centres. Liver Int 2008;28:821–7. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

