MGAT1 and Complex N-Glycans Regulate ERK Signaling During Spermatogenesis
- PMID: 29386567
- PMCID: PMC5792458
- DOI: 10.1038/s41598-018-20465-3
MGAT1 and Complex N-Glycans Regulate ERK Signaling During Spermatogenesis
Erratum in
-
Author Correction: MGAT1 and Complex N-Glycans Regulate ERK Signaling During Spermatogenesis.Sci Rep. 2023 Feb 28;13(1):3407. doi: 10.1038/s41598-023-30418-0. Sci Rep. 2023. PMID: 36854771 Free PMC article. No abstract available.
Abstract
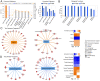
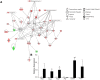
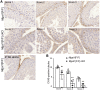
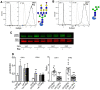
Mechanisms that regulate spermatogenesis in mice are important to define as they often apply to fertility in man. We previously showed that conditional deletion of the mouse Mgat1 gene (Mgat1 cKO) in spermatogonia causes a germ-cell autonomous defect leading to infertility. MGAT1 is the N-acetylglucosaminyltransferase (GlcNAcT-I) that initiates the synthesis of complex N-glycans. Mechanistic bases of MGAT1 loss were investigated in germ cells from 22- and 23-day males, before any changes in germ cell morphology were apparent. Gene expression changes induced by deletion of Mgat1 were determined using the Affymetrix gene chip Mouse Mogene 2.0 ST array, and relationships were investigated by bioinformatics including Gene Ontology (GO), Ingenuity Pathway Analysis (IPA), and Gene Set Enrichment Analysis (GSEA). The loss of complex N-glycans promoted the premature up-regulation of genes normally expressed later in spermatogenesis and spermiogenesis, and IPA and GSEA implicated ERK signaling. EGFR and PDGFRA transcripts and ERK1/2 signaling were reduced in 22-day Mgat1 cKO germ cells. Basigin, a germ cell target of MGAT1, activated ERK1/2 in CHO cells, but not in a Lec1 CHO mutant that lacks MGAT1 and complex N-glycans. Thus, MGAT1 is required to regulate ERK1/2 signaling during spermatogenesis, potentially via different mechanisms.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




Similar articles
-
Deletion of Mgat2 in spermatogonia blocks spermatogenesis.Front Cell Dev Biol. 2024 Sep 19;12:1428715. doi: 10.3389/fcell.2024.1428715. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 39364139 Free PMC article.
-
Complex N-glycans are essential, but core 1 and 2 mucin O-glycans, O-fucose glycans, and NOTCH1 are dispensable, for mammalian spermatogenesis.Biol Reprod. 2012 Jun 14;86(6):179. doi: 10.1095/biolreprod.111.098103. Print 2012 Jun. Biol Reprod. 2012. PMID: 22492969 Free PMC article.
-
Transgenic Rescue of Spermatogenesis in Males With Mgat1 Deleted in Germ Cells.Front Cell Dev Biol. 2020 Apr 2;8:212. doi: 10.3389/fcell.2020.00212. eCollection 2020. Front Cell Dev Biol. 2020. PMID: 32300591 Free PMC article.
-
Mgat1-dependent N-glycans are essential for the normal development of both vertebrate and invertebrate metazoans.Semin Cell Dev Biol. 2010 Aug;21(6):609-15. doi: 10.1016/j.semcdb.2010.02.010. Epub 2010 Mar 3. Semin Cell Dev Biol. 2010. PMID: 20206280 Review.
-
Hormonal regulation of spermatogenesis and spermiogenesis.J Steroid Biochem Mol Biol. 2008 Apr;109(3-5):323-30. doi: 10.1016/j.jsbmb.2008.03.004. Epub 2008 Mar 6. J Steroid Biochem Mol Biol. 2008. PMID: 18400489 Review.
Cited by
-
Genetic and genomic characterization of vulva size traits in Yorkshire and Landrace gilts.BMC Genet. 2020 Mar 12;21(1):28. doi: 10.1186/s12863-020-0834-9. BMC Genet. 2020. PMID: 32164558 Free PMC article.
-
Deletion of Mgat2 in spermatogonia blocks spermatogenesis.Front Cell Dev Biol. 2024 Sep 19;12:1428715. doi: 10.3389/fcell.2024.1428715. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 39364139 Free PMC article.
-
Global characterization of mouse testis O-glycoproteome landscape during spermatogenesis.Nat Commun. 2025 Mar 18;16(1):2676. doi: 10.1038/s41467-025-57980-7. Nat Commun. 2025. PMID: 40102425 Free PMC article.
-
Genetics of glycosylation in mammalian development and disease.Nat Rev Genet. 2024 Oct;25(10):715-729. doi: 10.1038/s41576-024-00725-x. Epub 2024 May 9. Nat Rev Genet. 2024. PMID: 38724711 Review.
-
The Central Role of Cadherins in Gonad Development, Reproduction, and Fertility.Int J Mol Sci. 2020 Nov 4;21(21):8264. doi: 10.3390/ijms21218264. Int J Mol Sci. 2020. PMID: 33158211 Free PMC article. Review.
References
-
- O’Bryan, M. K. & de Kretser, D. Mouse models for genes involved in impaired spermatogenesis. Int J Androl29, 76–89; discussion 105–108, 10.1111/j.1365-2605.2005.00614.x (2006). - PubMed
-
- Stanley, P. Taniguchi, N. & Aebi, M. In: Varki, A. et al. editors. Essentials of Glycobiology [Internet]. 3rd edition. Cold Spring Harbor (NY): Cold Spring Harbor Laboratory Press; Chapter 9, (2017).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

