A high HIV-1 strain variability in London, UK, revealed by full-genome analysis: Results from the ICONIC project
- PMID: 29389981
- PMCID: PMC5794160
- DOI: 10.1371/journal.pone.0192081
A high HIV-1 strain variability in London, UK, revealed by full-genome analysis: Results from the ICONIC project
Abstract
Background & methods: The ICONIC project has developed an automated high-throughput pipeline to generate HIV nearly full-length genomes (NFLG, i.e. from gag to nef) from next-generation sequencing (NGS) data. The pipeline was applied to 420 HIV samples collected at University College London Hospitals NHS Trust and Barts Health NHS Trust (London) and sequenced using an Illumina MiSeq at the Wellcome Trust Sanger Institute (Cambridge). Consensus genomes were generated and subtyped using COMET, and unique recombinants were studied with jpHMM and SimPlot. Maximum-likelihood phylogenetic trees were constructed using RAxML to identify transmission networks using the Cluster Picker.
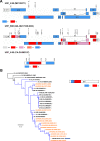
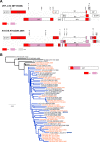
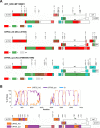
Results: The pipeline generated sequences of at least 1Kb of length (median = 7.46Kb, IQR = 4.01Kb) for 375 out of the 420 samples (89%), with 174 (46.4%) being NFLG. A total of 365 sequences (169 of them NFLG) corresponded to unique subjects and were included in the down-stream analyses. The most frequent HIV subtypes were B (n = 149, 40.8%) and C (n = 77, 21.1%) and the circulating recombinant form CRF02_AG (n = 32, 8.8%). We found 14 different CRFs (n = 66, 18.1%) and multiple URFs (n = 32, 8.8%) that involved recombination between 12 different subtypes/CRFs. The most frequent URFs were B/CRF01_AE (4 cases) and A1/D, B/C, and B/CRF02_AG (3 cases each). Most URFs (19/26, 73%) lacked breakpoints in the PR+RT pol region, rendering them undetectable if only that was sequenced. Twelve (37.5%) of the URFs could have emerged within the UK, whereas the rest were probably imported from sub-Saharan Africa, South East Asia and South America. For 2 URFs we found highly similar pol sequences circulating in the UK. We detected 31 phylogenetic clusters using the full dataset: 25 pairs (mostly subtypes B and C), 4 triplets and 2 quadruplets. Some of these were not consistent across different genes due to inter- and intra-subtype recombination. Clusters involved 70 sequences, 19.2% of the dataset.
Conclusions: The initial analysis of genome sequences detected substantial hidden variability in the London HIV epidemic. Analysing full genome sequences, as opposed to only PR+RT, identified previously undetected recombinants. It provided a more reliable description of CRFs (that would be otherwise misclassified) and transmission clusters.
Conflict of interest statement
Figures


References
-
- LANL, 2017. Los Alamos HIV database. Last accessed: May 1, 2017. Available from: http://www.hiv.lanl.gov.
-
- LANL, 2017. HIV Circulating Recombinant Forms (CRFs). Last accessed: May 1, 2017. Available from: http://www.hiv.lanl.gov/content/sequence/HIV/CRFs/CRFs.html.
-
- Hemelaar J, Gouws E, Ghys PD, Osmanov S. Global trends in molecular epidemiology of HIV-1 during 2000–2007. AIDS. 2011; 25: 679–89. doi: 10.1097/QAD.0b013e328342ff93 - DOI - PMC - PubMed
-
- Beloukas A, Psarris A, Giannelou P, Kostaki E, Hatzakis A, Paraskevis D. Molecular epidemiology of HIV-1 infection in Europe: An overview. Infect Genet Evol. 2016; 46: 180–9. doi: 10.1016/j.meegid.2016.06.033 - DOI - PubMed
-
- Dolling D, Hué S, Delpech V, Fearnhill E, Leigh-Brown A, Geretti AM, et al. The increasing genetic diversity of HIV-1 in the UK, 2002–2010. AIDS. 2014; 28: 773–80. doi: 10.1097/QAD.0000000000000119 - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

