Expression profile and promoter analysis of HEPIS
- PMID: 29399063
- PMCID: PMC5763650
- DOI: 10.3892/etm.2017.5374
Expression profile and promoter analysis of HEPIS
Abstract
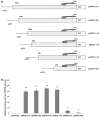
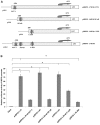
Human embryo lung cellular protein interacting with severe acute respiratory syndrome-coronavirus nonstructural protein-10 (HEPIS) is a novel transcriptional repressor, the expression profile and promoter activity of which have not been well studied. In the present study, in situ hybridization of RNA was used to study differential HEPIS expression levels in different types of cancer and normal tissues. A total of six truncated lengths of the HEPIS promoter regulatory sequences were cloned into the pGL3-basic vector, and reverse transcription-quantitative polymerase chain reaction (RT-qPCR) and dual luciferase reporter assays were performed. The results of RT-qPCR demonstrated that HEPIS expression levels differed across four breast cancer cell lines. The results of the dual luciferase reporter assays revealed that the activities of the reporter gene fragments spanning -1334/+373, -1203/+373, -1060/+373 and -899/+373 bp were higher compared with the reporter gene fragments spanning -759/+373 and -279/+373 bp. A search of the transcription factor database TRANSFAC identified numerous octamer transcription factor-1 (OCT-1), nuclear factor (NF)-κB and C-JUN transcription factor binding sites located on the HEPIS promoter (pHEPIS). Furthermore, the results revealed that mutations of the OCT-1 (-1236/-1223 bp), NF-κB (-1186/-1176 bp) and C-JUN (-856/-846 bp) sites on the human pHEPIS resulted in a decrease in luciferase activity. A chromatin immunoprecipitation assay revealed that OCT-1, NF-κB and C-JUN bound to pHEPIS in a site-dependent manner at the basal state. The TRANSFAC database was used to analyze the pHEPIS of multiple species and several activator protein-1, NF-κB and OCT-1 transcription factor binding sites were predicted. In conclusion, the results of the present study suggest that HEPIS is expressed at different levels in multiple organs and breast cancer cell lines. Furthermore, these findings indicate that OCT-1, NF-κB and C-JUN transcription factors are associated with transcriptional regulation of the HEPIS gene.
Keywords: core promoter; expression profile; human embryo lung cellular protein interacting with severe acute respiratory syndrome-coronavirus nonstructural protein-10; transcriptional regulation.
Figures




Similar articles
-
Expression and role of HEPIS in breast cancer.Oncol Lett. 2019 Dec;18(6):6648-6656. doi: 10.3892/ol.2019.10993. Epub 2019 Oct 17. Oncol Lett. 2019. PMID: 31788121 Free PMC article.
-
Identification of a novel transcriptional repressor (HEPIS) that interacts with nsp-10 of SARS coronavirus.Viral Immunol. 2008 Jun;21(2):153-62. doi: 10.1089/vim.2007.0108. Viral Immunol. 2008. PMID: 18433331
-
Regulation of the rat CYP4A2 gene promoter by c-Jun and octamer binding protein-1.Int J Biochem Cell Biol. 2007;39(6):1235-47. doi: 10.1016/j.biocel.2007.03.019. Epub 2007 Apr 1. Int J Biochem Cell Biol. 2007. PMID: 17481938
-
Oct-1 and nuclear factor Y bind to the SURG-1 element to direct basal and gonadotropin-releasing hormone (GnRH)-stimulated mouse GnRH receptor gene transcription.Mol Endocrinol. 2005 Jan;19(1):148-62. doi: 10.1210/me.2004-0025. Epub 2004 Sep 23. Mol Endocrinol. 2005. PMID: 15388790
-
NF-κB induces abnormal centrosome amplification by upregulation of CDK2 in laryngeal squamous cell cancer.Int J Oncol. 2011 Oct;39(4):915-24. doi: 10.3892/ijo.2011.1125. Epub 2011 Jul 15. Int J Oncol. 2011. PMID: 21769424
Cited by
-
Expression and role of HEPIS in breast cancer.Oncol Lett. 2019 Dec;18(6):6648-6656. doi: 10.3892/ol.2019.10993. Epub 2019 Oct 17. Oncol Lett. 2019. PMID: 31788121 Free PMC article.
References
-
- Sterling K, Bresnick E. Oct-1 transcription factor is a negative regulator of rat CYP1A1 expression via an octamer sequence in its negative regulatory element. Mol Pharmacol. 1996;49:329–337. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
