Cell fate in antiviral response arises in the crosstalk of IRF, NF-κB and JAK/STAT pathways
- PMID: 29402958
- PMCID: PMC5799375
- DOI: 10.1038/s41467-017-02640-8
Cell fate in antiviral response arises in the crosstalk of IRF, NF-κB and JAK/STAT pathways
Abstract
The innate immune system processes pathogen-induced signals into cell fate decisions. How information is turned to decision remains unknown. By combining stochastic mathematical modelling and experimentation, we demonstrate that feedback interactions between the IRF3, NF-κB and STAT pathways lead to switch-like responses to a viral analogue, poly(I:C), in contrast to pulse-like responses to bacterial LPS. Poly(I:C) activates both IRF3 and NF-κB, a requirement for induction of IFNβ expression. Autocrine IFNβ initiates a JAK/STAT-mediated positive-feedback stabilising nuclear IRF3 and NF-κB in first responder cells. Paracrine IFNβ, in turn, sensitises second responder cells through a JAK/STAT-mediated positive feedforward pathway that upregulates the positive-feedback components: RIG-I, PKR and OAS1A. In these sensitised cells, the 'live-or-die' decision phase following poly(I:C) exposure is shorter-they rapidly produce antiviral responses and commit to apoptosis. The interlinked positive feedback and feedforward signalling is key for coordinating cell fate decisions in cellular populations restricting pathogen spread.
Conflict of interest statement
The authors declare no competing financial interests.
Figures


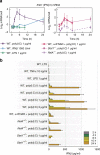





References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

