Effect of HIV/HCV Co-Infection on the Protease Evolution of HIV-1B: A Pilot Study in a Pediatric Population
- PMID: 29403002
- PMCID: PMC5799169
- DOI: 10.1038/s41598-018-19312-2
Effect of HIV/HCV Co-Infection on the Protease Evolution of HIV-1B: A Pilot Study in a Pediatric Population
Abstract
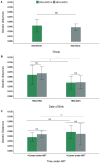
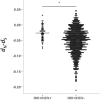
This pilot study evaluates in pediatric patients the impact of HIV/HCV coinfection in the molecular evolution of the HIV-1 subtype B protease (HIV-1BPR). For this study, HIV-1B/HCV coinfected (15) and HIV-1B monoinfected (56) patients with available HIV-1B pol sequences were enrolled. Both groups of patients had comparable gender frequencies and average age, time of infection, antiretroviral treatment (ART) exposure and time under ART. Prevalence of drug resistance mutations (DRM), genetic diversity, number of synonymous (dS) and non-synonymous (dN) mutations per site and selection pressures (dN - dS) in the HIV-1BPR were estimated and compared between mono- and coinfected patients. Both HIV-1B populations presented similar genetic diversity (0.050 ± 0.02 vs. 0.045 ± 0.01) and dS (0.074 ± 0.03 vs. 0.078 ± 0.04). In turn, in coinfected patients the HIV-1BPR had higher dN (0.045 ± 0.01 vs. 0.024 ± 0.01) and dN-dS (-0.026 ± 0.02 vs. -0.048 ± 0.04) values, and less amino acid sites under purifying selection (4.2% vs. 42.1%) than in monoinfected patients. Accordingly, in co-infection with HCV, the HIV-1BPR sites 50, 53, 82, 84 and 88 - associated with resistance to PIs - were under neutral evolution, whereas these sites were under purifying selection in monoinfected patients. This pilot study suggests that HIV-1B may evolve differently in the presence than in the absence of HCV.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
Hepatitis C virus (HCV) protease variability and anti-HCV protease inhibitor resistance in HIV/HCV-coinfected patients.HIV Med. 2011 Sep;12(8):506-9. doi: 10.1111/j.1468-1293.2011.00913.x. Epub 2011 Mar 16. HIV Med. 2011. PMID: 21410862
-
Resistance-associated mutations to HCV protease inhibitors naturally pre-existed in HIV/HCV coinfected, treatment-naïve patients.Clin Res Hepatol Gastroenterol. 2016 Nov;40(5):597-604. doi: 10.1016/j.clinre.2016.02.004. Epub 2016 Mar 23. Clin Res Hepatol Gastroenterol. 2016. PMID: 27016893
-
Resistance mutations are rare among protease inhibitor treatment-naive hepatitis C genotype-1 patients with or without HIV coinfection.Antivir Ther. 2015;20(3):281-7. doi: 10.3851/IMP2873. Epub 2014 Oct 3. Antivir Ther. 2015. PMID: 25279715
-
Impact of Clinical Parameters in the Intrahost Evolution of HIV-1 Subtype B in Pediatric Patients: A Machine Learning Approach.Genome Biol Evol. 2017 Oct 1;9(10):2715-2726. doi: 10.1093/gbe/evx193. Genome Biol Evol. 2017. PMID: 29044435 Free PMC article.
-
Treatment of chronic HCV genotype 1 coinfection.Curr HIV/AIDS Rep. 2015 Sep;12(3):326-35. doi: 10.1007/s11904-015-0278-4. Curr HIV/AIDS Rep. 2015. PMID: 26228050 Review.
Cited by
-
Drug Resistance Prediction Using Deep Learning Techniques on HIV-1 Sequence Data.Viruses. 2020 May 19;12(5):560. doi: 10.3390/v12050560. Viruses. 2020. PMID: 32438586 Free PMC article.
References
-
- Alizon S, de Roode JC, Michalakis Y. Multiple infections and the evolution of virulence. In: Drake J, editor. Ecol Lett. 2013. pp. 556–67. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

