Integrative analysis of methylomic and transcriptomic data in fetal sheep muscle tissues in response to maternal diet during pregnancy
- PMID: 29409445
- PMCID: PMC5801776
- DOI: 10.1186/s12864-018-4509-0
Integrative analysis of methylomic and transcriptomic data in fetal sheep muscle tissues in response to maternal diet during pregnancy
Abstract
Background: Numerous studies have established a link between maternal diet and the physiological and metabolic phenotypes of their offspring. In previous studies in sheep, we demonstrated that different maternal diets altered the transcriptome of fetal tissues. However, the mechanisms underlying transcriptomic changes are poorly understood. DNA methylation is an epigenetic mark regulating transcription and is largely influenced by dietary components of the one-carbon cycle that generate the methyl group donor, SAM. Therefore, in the present study, we tested whether different maternal diets during pregnancy would alter the DNA methylation and gene expression patterns in fetal tissues.
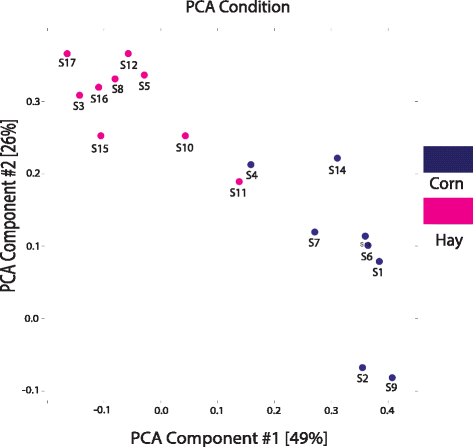
Results: Pregnant ewes were randomly divided into two groups which received either hay or corn diet from mid-gestation (day 67 ± 5) until day 131 ± 1 when fetuses were collected by necropsy. A total of 1516 fetal longissimus dorsi (LD) tissues were used for DNA methylation analysis and gene expression profiling. Whole genome DNA methylation using methyl-binding domain enrichment analysis revealed 60 differentially methylated regions (DMRs) between hay and corn fetuses with 39 DMRs more highly methylated in the hay fetuses vs. 21 DMRs more highly methylated in the corn fetuses. Three DMRs (LPAR3, PLIN5-PLIN4, and the differential methylation of a novel lincRNA) were validated using bisulfite sequencing. These DMRs were associated with differential gene expression. Additionally, significant DNA methylation differences were found at the single CpG level. Integrative methylome and transcriptome analysis revealed an association between gene expression and inter-/intragenic methylated regions. Furthermore, intragenic DMRs were found to be associated with expression of neighboring genes.
Conclusions: The findings of this study imply that maternal diet from mid- to late-gestation can shape the epigenome and the transcriptome of fetal tissues, and putatively affect phenotypes of the lambs.
Keywords: DNA methylation; Differentially-methylated region; Gene expression; Maternal diet.
Conflict of interest statement
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Figures



References
-
- Jaenisch R, Bird A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat. Genet. 2003;33 Suppl:245–254. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

