Asparagine bioavailability governs metastasis in a model of breast cancer
- PMID: 29414946
- PMCID: PMC5898613
- DOI: 10.1038/nature25465
Asparagine bioavailability governs metastasis in a model of breast cancer
Erratum in
-
Erratum: Asparagine bioavailability governs metastasis in a model of breast cancer.Nature. 2018 Apr 4;556(7699):135. doi: 10.1038/nature26162. Nature. 2018. PMID: 29620732
Abstract
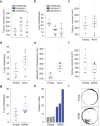
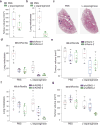
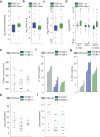
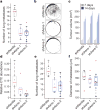
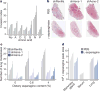
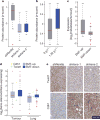
Using a functional model of breast cancer heterogeneity, we previously showed that clonal sub-populations proficient at generating circulating tumour cells were not all equally capable of forming metastases at secondary sites. A combination of differential expression and focused in vitro and in vivo RNA interference screens revealed candidate drivers of metastasis that discriminated metastatic clones. Among these, asparagine synthetase expression in a patient's primary tumour was most strongly correlated with later metastatic relapse. Here we show that asparagine bioavailability strongly influences metastatic potential. Limiting asparagine by knockdown of asparagine synthetase, treatment with l-asparaginase, or dietary asparagine restriction reduces metastasis without affecting growth of the primary tumour, whereas increased dietary asparagine or enforced asparagine synthetase expression promotes metastatic progression. Altering asparagine availability in vitro strongly influences invasive potential, which is correlated with an effect on proteins that promote the epithelial-to-mesenchymal transition. This provides at least one potential mechanism for how the bioavailability of a single amino acid could regulate metastatic progression.
Conflict of interest statement
Figures









Comment in
-
Asparagine Bioavailability Drives Breast Cancer Metastasis.Cancer Discov. 2018 Apr;8(4):381. doi: 10.1158/2159-8290.CD-RW2018-026. Epub 2018 Feb 16. Cancer Discov. 2018. PMID: 29453237
-
Asparagine and Glutamine: Co-conspirators Fueling Metastasis.Cell Metab. 2018 May 1;27(5):947-949. doi: 10.1016/j.cmet.2018.04.012. Cell Metab. 2018. PMID: 29719230
References
-
- Miller FR, Miller BE, Heppner GH. Characterization of metastatic heterogeneity among subpopulations of a single mouse mammary tumor: heterogeneity in phenotypic stability. Invasion Metastasis. 1983;3:22–31. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

