The Mouse Cortical Connectome, Characterized by an Ultra-Dense Cortical Graph, Maintains Specificity by Distinct Connectivity Profiles
- PMID: 29420935
- PMCID: PMC5958229
- DOI: 10.1016/j.neuron.2017.12.037
The Mouse Cortical Connectome, Characterized by an Ultra-Dense Cortical Graph, Maintains Specificity by Distinct Connectivity Profiles
Abstract
The inter-areal wiring pattern of the mouse cerebral cortex was analyzed in relation to a refined parcellation of cortical areas. Twenty-seven retrograde tracer injections were made in 19 areas of a 47-area parcellation of the mouse neocortex. Flat mounts of the cortex and multiple histological markers enabled detailed counts of labeled neurons in individual areas. The observed log-normal distribution of connection weights to each cortical area spans 5 orders of magnitude and reveals a distinct connectivity profile for each area, analogous to that observed in macaques. The cortical network has a density of 97%, considerably higher than the 66% density reported in macaques. A weighted graph analysis reveals a similar global efficiency but weaker spatial clustering compared with that reported in macaques. The consistency, precision of the connectivity profile, density, and weighted graph analysis of the present data differ significantly from those obtained in earlier studies in the mouse.
Keywords: anatomy; connectivity; log-normal; neocortex; retrograde; rodent; tract-tracing.
Copyright © 2017 Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors declare no competing interests.
Figures

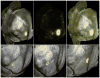

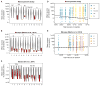
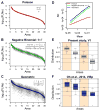
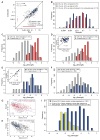
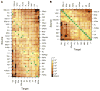

References
-
- Akaike H. A new look at the statistical model identification. IEEE Transactions on Automatic Control. 1974;19:716–723.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

