Polymorphisms of long non-coding RNA HOTAIR with breast cancer susceptibility and clinical outcomes for a southeast Chinese Han population
- PMID: 29423075
- PMCID: PMC5790492
- DOI: 10.18632/oncotarget.23343
Polymorphisms of long non-coding RNA HOTAIR with breast cancer susceptibility and clinical outcomes for a southeast Chinese Han population
Abstract
Hox transcript antisense intergenic RNA (HOTAIR) is a well-known long non-coding RNA (lncRNA) which participates in tumorigenesis and progress of multiple cancers. However, the associations among polymorphisms on HOTAIR, breast cancer (BC) susceptibility and clinical outcomes have remained obscure. In this case-control study, we assessed the interaction between three lncRNA HOTAIR single nucleotide polymorphisms (SNPs) (rs1899663, rs4759314 and rs7958904) on the risk and clinical outcome of breast cancer in a Chinese Han population. In total, 969 breast cancer cases and 970 healthy controls were enrolled in this study. Associations among genotypes, BC risk and survival were evaluated by univariate and multivariate logistic regression to estimate the odds ratio (OR), hazard ratio (HR) and its 95% confidence interval (CI). The disease-free survival (DFS) and overall survival (OS) was calculated by the Kaplan-Meier method. We found that the T allele of rs1899663 and C allele of rs7958904 both achieved significant differences between cases and controls in the single locus analyses (P = 0.017 and 0.010, respectively). Multivariate analyses also revealed the rs1899663 TT genotype and rs7958904 CC genotype were both at higher risk of breast cancer compared with the GG homozygotes (OR = 2.08, 95% CI = 1.20-3.60 and OR = 1.45, 95% CI = 1.01-2.08, respectively). In survival analysis, we observed that the T allele of rs1899663 presented significant differences for both DFS (HR = 1.64, 95% CI = 1.12-2.40) and OS (HR = 2.10, 95% CI = 1.29-3.42) in younger subjects (age ≤ 40). Our findings may provide new insights into the associations among the genetic susceptibility, the fine classifications and the prognosis of breast cancer. Further studies with larger sample size and functional research should also be conducted to validate our findings and better elucidate the underlying biological mechanisms.
Keywords: HOTAIR; breast cancer; genetic susceptibility; polymorphisms; prognosis.
Conflict of interest statement
CONFLICTS OF INTEREST The authors have declared that no competing interests exist.
Figures
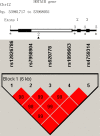
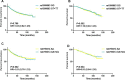

References
-
- Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108. https://doi.org/10.3322/caac.21262. - DOI - PubMed
-
- Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017;67:7–30. https://doi.org/10.3322/caac.21387. - DOI - PubMed
-
- Fan L, Strasser-Weippl K, Li JJ, St Louis J, Finkelstein DM, Yu KD, Chen WQ, Shao ZM, Goss PE. Breast cancer in China. Lancet Oncol. 2014;15:e279–89. https://doi.org/10.1016/S1470-2045(13)70567-9. - DOI - PubMed
-
- Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66:115–32. https://doi.org/10.3322/caac.21338. - DOI - PubMed
-
- Key TJ, Verkasalo PK, Banks E. Epidemiology of breast cancer. Lancet Oncol. 2001;2:133–40. https://doi.org/10.1016/S1470-2045(00)00254-0. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

