Loss of microRNA-128 promotes cardiomyocyte proliferation and heart regeneration
- PMID: 29453456
- PMCID: PMC5816015
- DOI: 10.1038/s41467-018-03019-z
Loss of microRNA-128 promotes cardiomyocyte proliferation and heart regeneration
Abstract
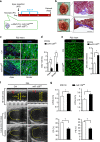
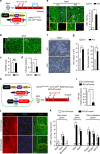
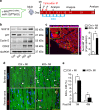
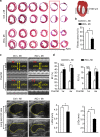
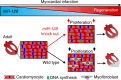
The goal of replenishing the cardiomyocyte (CM) population using regenerative therapies following myocardial infarction (MI) is hampered by the limited regeneration capacity of adult CMs, partially due to their withdrawal from the cell cycle. Here, we show that microRNA-128 (miR-128) is upregulated in CMs during the postnatal switch from proliferation to terminal differentiation. In neonatal mice, cardiac-specific overexpression of miR-128 impairs CM proliferation and cardiac function, while miR-128 deletion extends proliferation of postnatal CMs by enhancing expression of the chromatin modifier SUZ12, which suppresses p27 (cyclin-dependent kinase inhibitor) expression and activates the positive cell cycle regulators Cyclin E and CDK2. Furthermore, deletion of miR-128 promotes cell cycle re-entry of adult CMs, thereby reducing the levels of fibrosis, and attenuating cardiac dysfunction in response to MI. These results suggest that miR-128 serves as a critical regulator of endogenous CM proliferation, and might be a novel therapeutic target for heart repair.
Conflict of interest statement
The authors declare no competing financial interests.
Figures
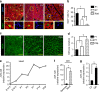

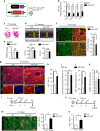
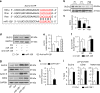
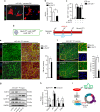
Similar articles
-
MicroRNA-34a Plays a Key Role in Cardiac Repair and Regeneration Following Myocardial Infarction.Circ Res. 2015 Aug 14;117(5):450-9. doi: 10.1161/CIRCRESAHA.117.305962. Epub 2015 Jun 16. Circ Res. 2015. PMID: 26082557 Free PMC article.
-
Melatonin promotes cardiomyocyte proliferation and heart repair in mice with myocardial infarction via miR-143-3p/Yap/Ctnnd1 signaling pathway.Acta Pharmacol Sin. 2021 Jun;42(6):921-931. doi: 10.1038/s41401-020-0495-2. Epub 2020 Aug 24. Acta Pharmacol Sin. 2021. PMID: 32839503 Free PMC article.
-
MicroRNA-210-mediated proliferation, survival, and angiogenesis promote cardiac repair post myocardial infarction in rodents.J Mol Med (Berl). 2017 Dec;95(12):1369-1385. doi: 10.1007/s00109-017-1591-8. Epub 2017 Sep 25. J Mol Med (Berl). 2017. PMID: 28948298 Free PMC article.
-
Recent Insights into Endogenous Mammalian Cardiac Regeneration Post-Myocardial Infarction.Int J Mol Sci. 2024 Nov 1;25(21):11747. doi: 10.3390/ijms252111747. Int J Mol Sci. 2024. PMID: 39519298 Free PMC article. Review.
-
The role of microRNAs in cardiac development and regenerative capacity.Am J Physiol Heart Circ Physiol. 2016 Mar 1;310(5):H528-41. doi: 10.1152/ajpheart.00181.2015. Epub 2015 Dec 23. Am J Physiol Heart Circ Physiol. 2016. PMID: 26702142 Free PMC article. Review.
Cited by
-
Non-coding RNAs in Cardiac Regeneration.Front Physiol. 2021 Mar 24;12:650566. doi: 10.3389/fphys.2021.650566. eCollection 2021. Front Physiol. 2021. PMID: 33841185 Free PMC article. Review.
-
Ligand-Dependent Downregulation of Guanylyl Cyclase/Natriuretic Peptide Receptor-A: Role of miR-128 and miR-195.Int J Mol Sci. 2022 Nov 2;23(21):13381. doi: 10.3390/ijms232113381. Int J Mol Sci. 2022. PMID: 36362173 Free PMC article.
-
Return of the Tbx5; lineage-tracing reveals ventricular cardiomyocyte-like precursors in the injured adult mammalian heart.NPJ Regen Med. 2023 Mar 3;8(1):13. doi: 10.1038/s41536-023-00280-9. NPJ Regen Med. 2023. PMID: 36869039 Free PMC article.
-
Applications of miRNAs in cardiac development, disease progression and regeneration.Stem Cell Res Ther. 2019 Nov 21;10(1):336. doi: 10.1186/s13287-019-1451-2. Stem Cell Res Ther. 2019. PMID: 31752983 Free PMC article. Review.
-
Exercise-Induced Adult Cardiomyocyte Proliferation in Mammals.Front Physiol. 2021 Aug 30;12:729364. doi: 10.3389/fphys.2021.729364. eCollection 2021. Front Physiol. 2021. PMID: 34526914 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

