RaMP: A Comprehensive Relational Database of Metabolomics Pathways for Pathway Enrichment Analysis of Genes and Metabolites
- PMID: 29470400
- PMCID: PMC5876005
- DOI: 10.3390/metabo8010016
RaMP: A Comprehensive Relational Database of Metabolomics Pathways for Pathway Enrichment Analysis of Genes and Metabolites
Abstract
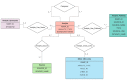
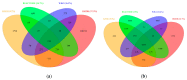
The value of metabolomics in translational research is undeniable, and metabolomics data are increasingly generated in large cohorts. The functional interpretation of disease-associated metabolites though is difficult, and the biological mechanisms that underlie cell type or disease-specific metabolomics profiles are oftentimes unknown. To help fully exploit metabolomics data and to aid in its interpretation, analysis of metabolomics data with other complementary omics data, including transcriptomics, is helpful. To facilitate such analyses at a pathway level, we have developed RaMP (Relational database of Metabolomics Pathways), which combines biological pathways from the Kyoto Encyclopedia of Genes and Genomes (KEGG), Reactome, WikiPathways, and the Human Metabolome DataBase (HMDB). To the best of our knowledge, an off-the-shelf, public database that maps genes and metabolites to biochemical/disease pathways and can readily be integrated into other existing software is currently lacking. For consistent and comprehensive analysis, RaMP enables batch and complex queries (e.g., list all metabolites involved in glycolysis and lung cancer), can readily be integrated into pathway analysis tools, and supports pathway overrepresentation analysis given a list of genes and/or metabolites of interest. For usability, we have developed a RaMP R package (https://github.com/Mathelab/RaMP-DB), including a user-friendly RShiny web application, that supports basic simple and batch queries, pathway overrepresentation analysis given a list of genes or metabolites of interest, and network visualization of gene-metabolite relationships. The package also includes the raw database file (mysql dump), thereby providing a stand-alone downloadable framework for public use and integration with other tools. In addition, the Python code needed to recreate the database on another system is also publicly available (https://github.com/Mathelab/RaMP-BackEnd). Updates for databases in RaMP will be checked multiple times a year and RaMP will be updated accordingly.
Keywords: metabolomics; pathway analysis; pathway database; transcriptomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Integration of Metabolomics and Transcriptomics to Identify Gene-Metabolite Relationships Specific to Phenotype.Methods Mol Biol. 2019;1928:441-468. doi: 10.1007/978-1-4939-9027-6_23. Methods Mol Biol. 2019. PMID: 30725469
-
RaMP-DB 2.0: a renovated knowledgebase for deriving biological and chemical insight from metabolites, proteins, and genes.Bioinformatics. 2023 Jan 1;39(1):btac726. doi: 10.1093/bioinformatics/btac726. Bioinformatics. 2023. PMID: 36373969 Free PMC article.
-
IntLIM: integration using linear models of metabolomics and gene expression data.BMC Bioinformatics. 2018 Mar 5;19(1):81. doi: 10.1186/s12859-018-2085-6. BMC Bioinformatics. 2018. PMID: 29506475 Free PMC article.
-
Metabolomics in severe traumatic brain injury: a scoping review.BMC Neurosci. 2023 Oct 16;24(1):54. doi: 10.1186/s12868-023-00824-1. BMC Neurosci. 2023. PMID: 37845610 Free PMC article.
-
The interactions and biological pathways among metabolomics products of patients with coronary heart disease.Biomed Pharmacother. 2024 Apr;173:116305. doi: 10.1016/j.biopha.2024.116305. Epub 2024 Feb 29. Biomed Pharmacother. 2024. PMID: 38422653 Review.
Cited by
-
Survey for Computer-Aided Tools and Databases in Metabolomics.Metabolites. 2022 Oct 21;12(10):1002. doi: 10.3390/metabo12101002. Metabolites. 2022. PMID: 36295904 Free PMC article. Review.
-
The metaRbolomics Toolbox in Bioconductor and beyond.Metabolites. 2019 Sep 23;9(10):200. doi: 10.3390/metabo9100200. Metabolites. 2019. PMID: 31548506 Free PMC article. Review.
-
The Omics Revolution Continues: The Maturation of High-Throughput Biological Data Sources.Yearb Med Inform. 2018 Aug;27(1):211-222. doi: 10.1055/s-0038-1667085. Epub 2018 Aug 29. Yearb Med Inform. 2018. PMID: 30157526 Free PMC article. Review.
-
PathMe: merging and exploring mechanistic pathway knowledge.BMC Bioinformatics. 2019 May 15;20(1):243. doi: 10.1186/s12859-019-2863-9. BMC Bioinformatics. 2019. PMID: 31092193 Free PMC article.
-
IntLIM 2.0: identifying multi-omic relationships dependent on discrete or continuous phenotypic measurements.Bioinform Adv. 2023 Feb 1;3(1):vbad009. doi: 10.1093/bioadv/vbad009. eCollection 2023. Bioinform Adv. 2023. PMID: 36922980 Free PMC article.
References
-
- Mathé E.A., Patterson A.D., Haznadar M., Manna S.K., Krausz K.W., Bowman E.D., Shields P.G., Idle J.R., Smith P.B., Anami K. Noninvasive urinary metabolomic profiling identifies diagnostic and prognostic markers in lung cancer. Cancer Res. 2014;74:3259–3270. doi: 10.1158/0008-5472.CAN-14-0109. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

