A Genome-Wide Association Study for Host Resistance to Ostreid Herpesvirus in Pacific Oysters (Crassostrea gigas)
- PMID: 29472307
- PMCID: PMC5873916
- DOI: 10.1534/g3.118.200113
A Genome-Wide Association Study for Host Resistance to Ostreid Herpesvirus in Pacific Oysters (Crassostrea gigas)
Abstract
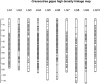
Ostreid herpesvirus (OsHV) can cause mass mortality events in Pacific oyster aquaculture. While various factors impact on the severity of outbreaks, it is clear that genetic resistance of the host is an important determinant of mortality levels. This raises the possibility of selective breeding strategies to improve the genetic resistance of farmed oyster stocks, thereby contributing to disease control. Traditional selective breeding can be augmented by use of genetic markers, either via marker-assisted or genomic selection. The aim of the current study was to investigate the genetic architecture of resistance to OsHV in Pacific oyster, to identify genomic regions containing putative resistance genes, and to inform the use of genomics to enhance efforts to breed for resistance. To achieve this, a population of ∼1,000 juvenile oysters were experimentally challenged with a virulent form of OsHV, with samples taken from mortalities and survivors for genotyping and qPCR measurement of viral load. The samples were genotyped using a recently-developed SNP array, and the genotype data were used to reconstruct the pedigree. Using these pedigree and genotype data, the first high density linkage map was constructed for Pacific oyster, containing 20,353 SNPs mapped to the ten pairs of chromosomes. Genetic parameters for resistance to OsHV were estimated, indicating a significant but low heritability for the binary trait of survival and also for viral load measures (h2 0.12 - 0.25). A genome-wide association study highlighted a region of linkage group 6 containing a significant QTL affecting host resistance. These results are an important step toward identification of genes underlying resistance to OsHV in oyster, and a step toward applying genomic data to enhance selective breeding for disease resistance in oyster aquaculture.
Keywords: GWAS; OsHV-1; SNP array; linkage map; oysters.
Copyright © 2018 Gutierrez et al.
Figures

References
-
- Azéma P., Lamy J.-B., Boudry P., Renault T., Travers M.-A., et al. , 2017. Genetic parameters of resistance to Vibrio aestuarianus, and OsHV-1 infections in the Pacific oyster, Crassostrea gigas, at three different life stages. Genet. Sel. Evol. 49(1): 23 10.1186/s12711-017-0297-2 - DOI - PMC - PubMed
-
- Bishop D. T., Cannings C., Skolnick M., Williamson J. A., Weir B. S., 1983. The number of polymorphic DNA clones required to map the human genome. Statistical Analysis of DNA Sequence Data, B.S. Weir, New York.
-
- Camara M. D., Yen S., Kaspar H. F., Kesarcodi-Watson A., King N., et al. , 2017. Assessment of heat shock and laboratory virus challenges to selectively breed for ostreid herpesvirus 1 (OsHV-1) resistance in the Pacific oyster, Crassostrea gigas. Aquaculture 469: 50–58. 10.1016/j.aquaculture.2016.11.031 - DOI
Publication types
MeSH terms
Substances
Grants and funding
- BBS/E/D/20211553/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/P013740/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/P013759/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/M026140/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- MR/K001744/1/MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
